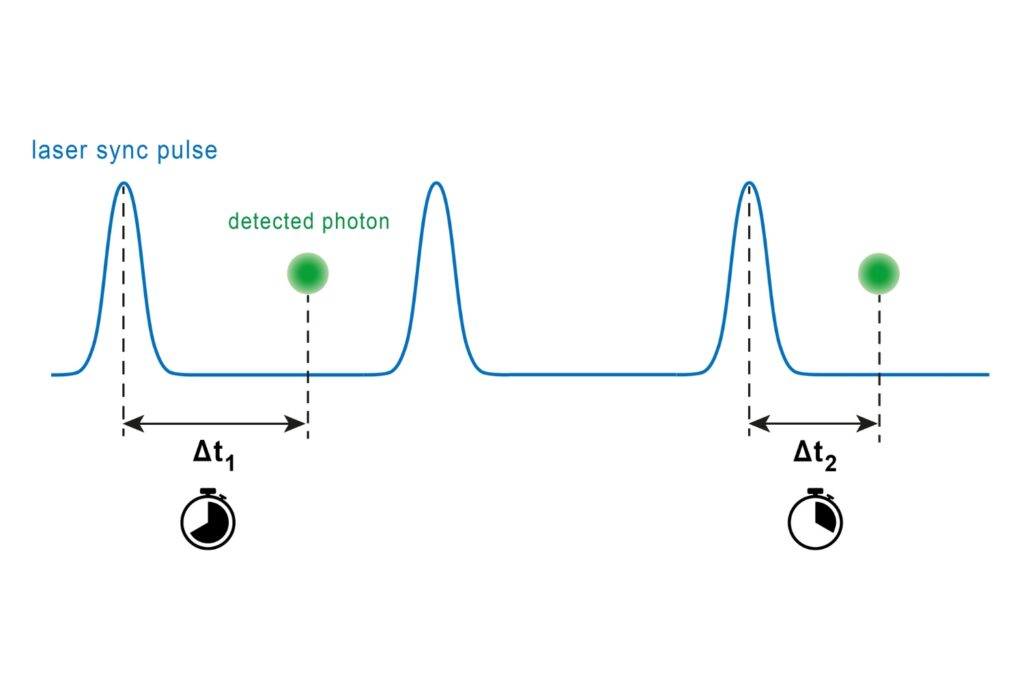
Scientific Applications and Methods
For Life Science, Materials Science, and Quantum Optics
Scientific discovery requires both the right research context and the appropriate measurement approach. Explore how different fields and techniques work together to reveal dynamic processes with high precision.
Find Your Research Focus
Explore Key Scientific Fields
Select a research field and switch between applications and methods to explore how measurement approaches enable insight into complex systems across disciplines.
Choose a research field*

Materials Science
Materials Science

Materials Science
Materials Science
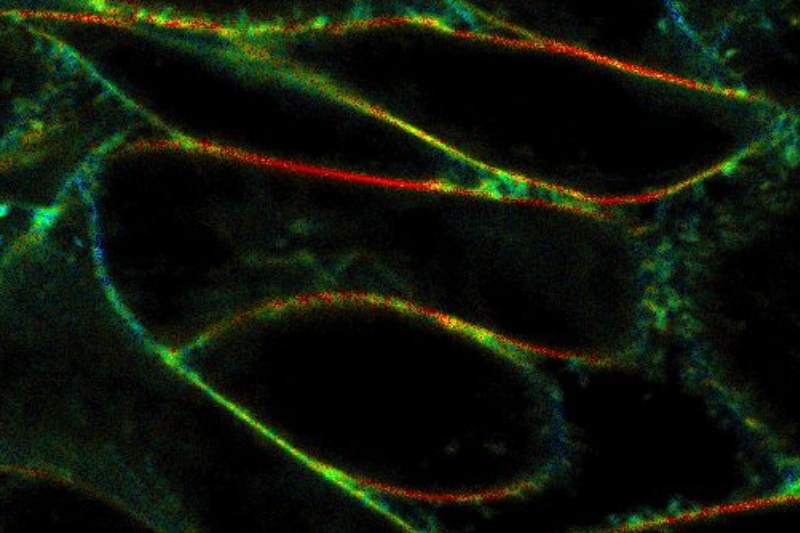
Life Science
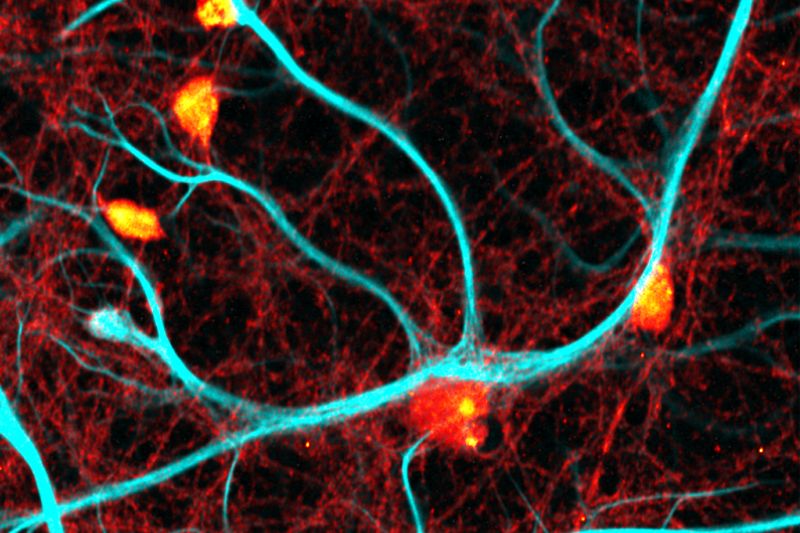
Life Science
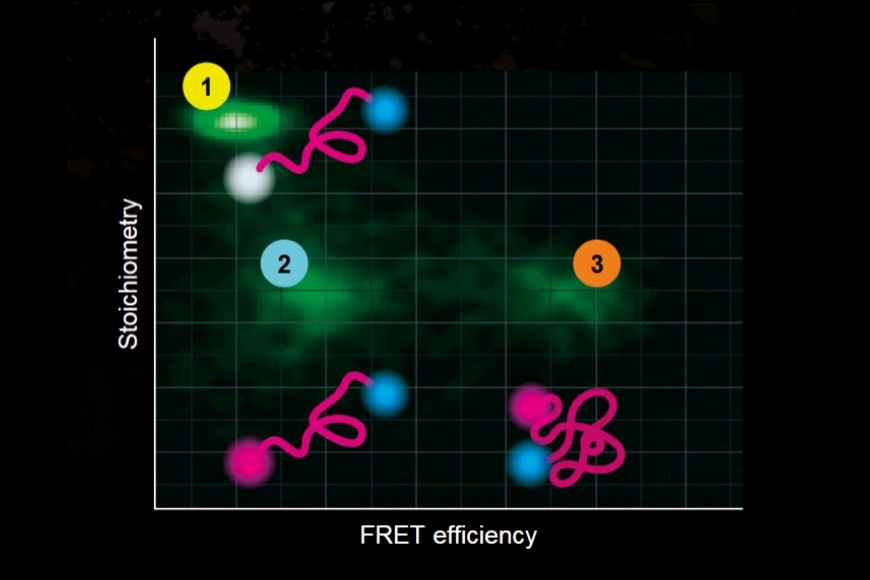
Life Science
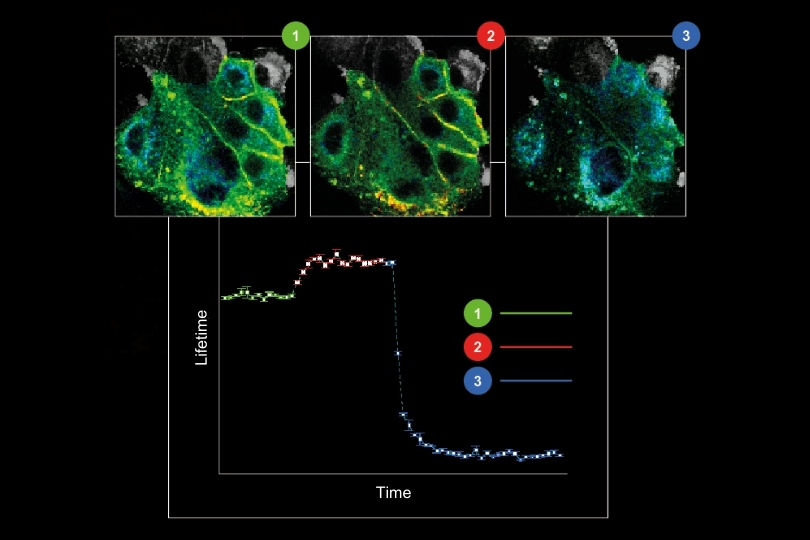
Life Science | Materials Science | Metrology
Materials Science
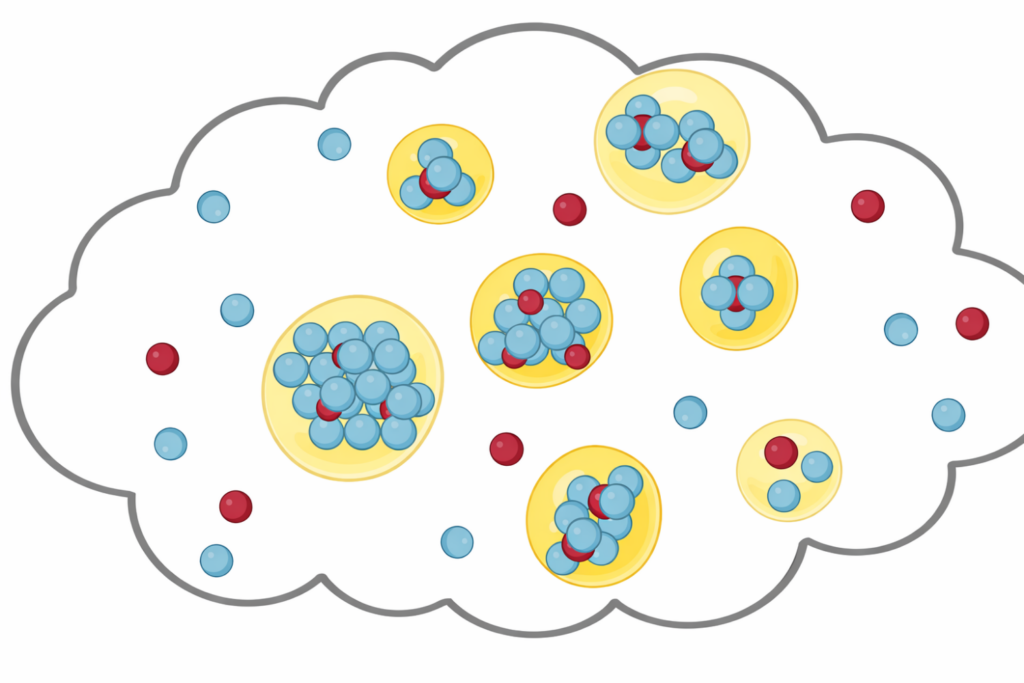
Life Science
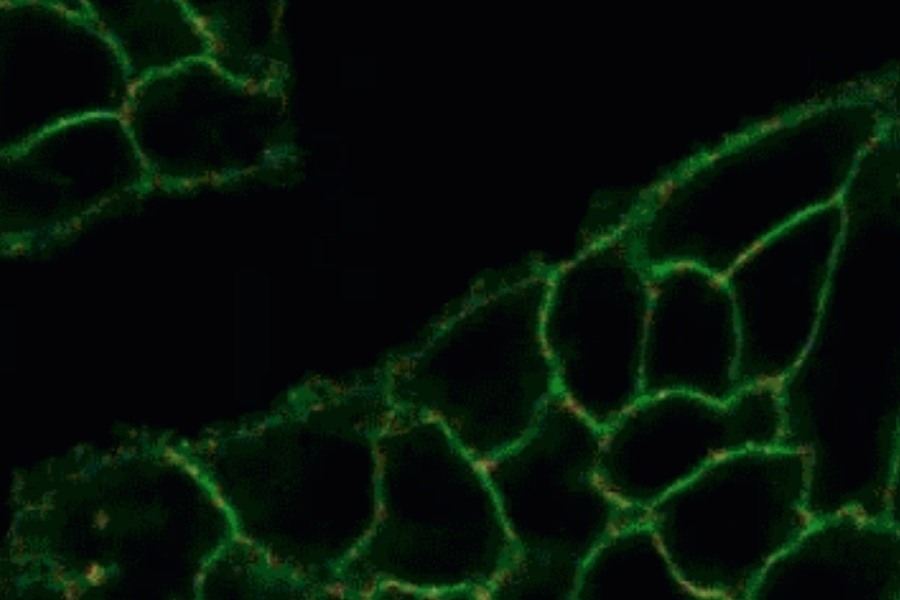
Life Science
Materials Science