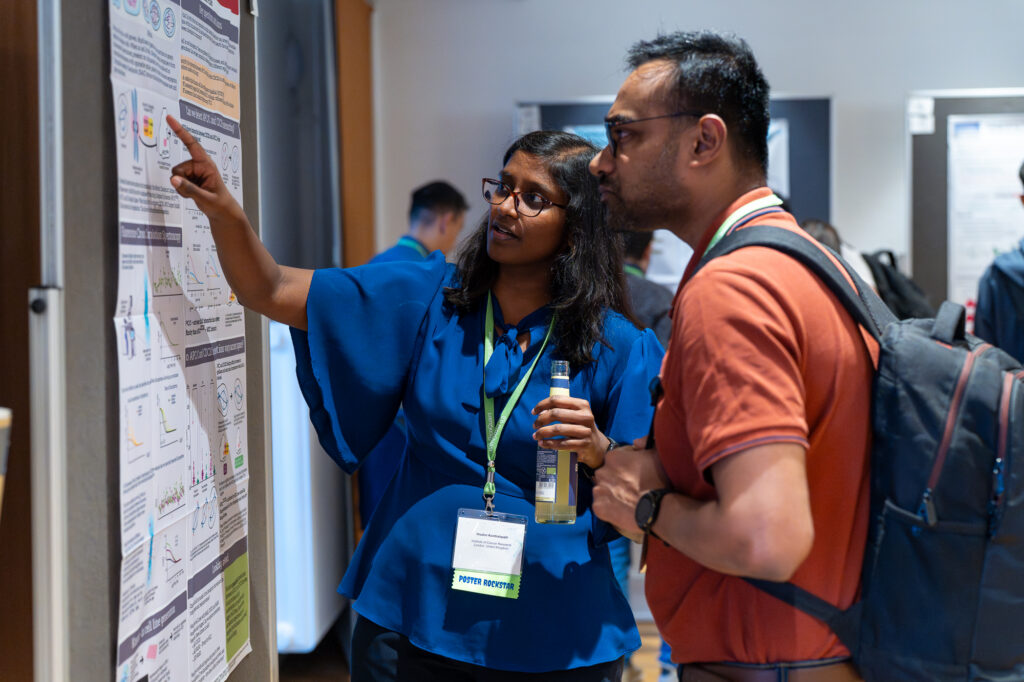
Be part of an exciting and stimulating conference where you can give a talk, present a poster, or attend without presenting. As always, we will award a “Best Student Talk” prize worth 750 Euro.
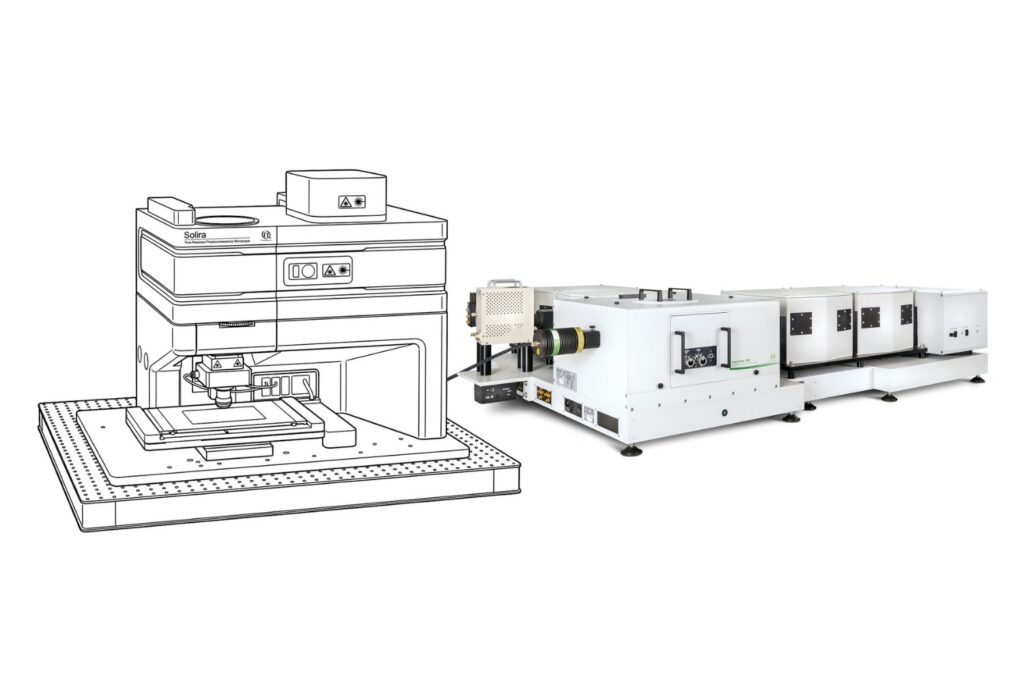
Immerse yourself in a groundbreaking exploration of ultra-sensitive optical detection down to the single molecule level as well as beyond the classical diffraction limit. PicoQuant’s renowned workshop provides an unparalleled platform for interdisciplinary collaboration and the exchange of cutting-edge research.
Discover a vast array of applications and methods in single molecule spectroscopy and advanced microscopy, presented through captivating talks and posters. Network with leading experts and like-minded researchers at our welcoming reception and workshop dinner.
Don’t miss this opportunity to be at the forefront of scientific innovation!
Covered topics include
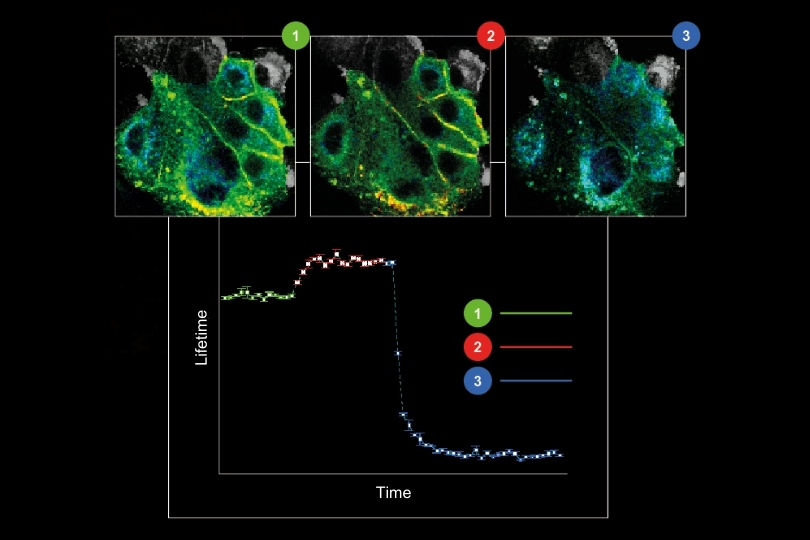
- Fluorescence Lifetime Imaging (FLIM)
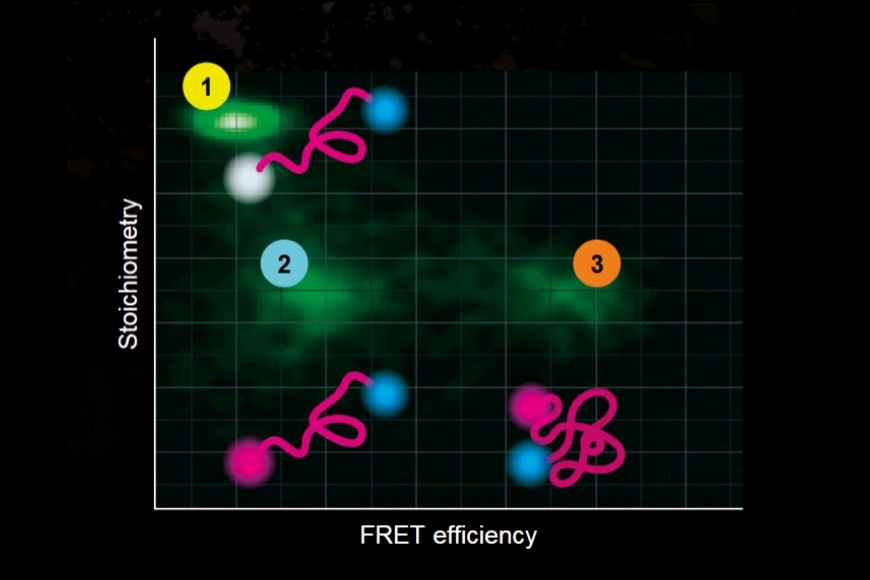
- Single molecule Förster Resonance Energy Transfer (smFRET)
- Polarization and Anisotropy based techniques
- Quantitative imaging methods
- New fluorescence sensors and labeling schemes
- Fluorescence Correlation Spectroscopy (FCS) and Fluorescence Lifetime Correlation Spectroscopy (FLCS)
- Image Scanning Microscopy (ISM)
- Single Molecule Localization Microscopy (e.g., PALM, STORM, dSTORM, GSDIM, PAINT)
- Open source data analysis solutions
- Big Data and Machine Learning approaches to superresolution and single molecule techniques
Both widefield and confocal fluorescence microscopy techniques are covered as well as in vivo and in vitro measurements with single molecule sensitivity.
Important Dates
Deadline for early bird registration: March 30, 2026
Deadline for abstract submission: June 8, 2026
Deadline for fee waiver application: May 11, 2026
Final deadline for workshop registration: August 31, 2026
Program available: End of August 2026