Products
- Microscope Systems
Most popular products

Solira
Simplify your materials characterization with one flexible TRPL microscope enabling multiple methods for precise and efficient analysis.
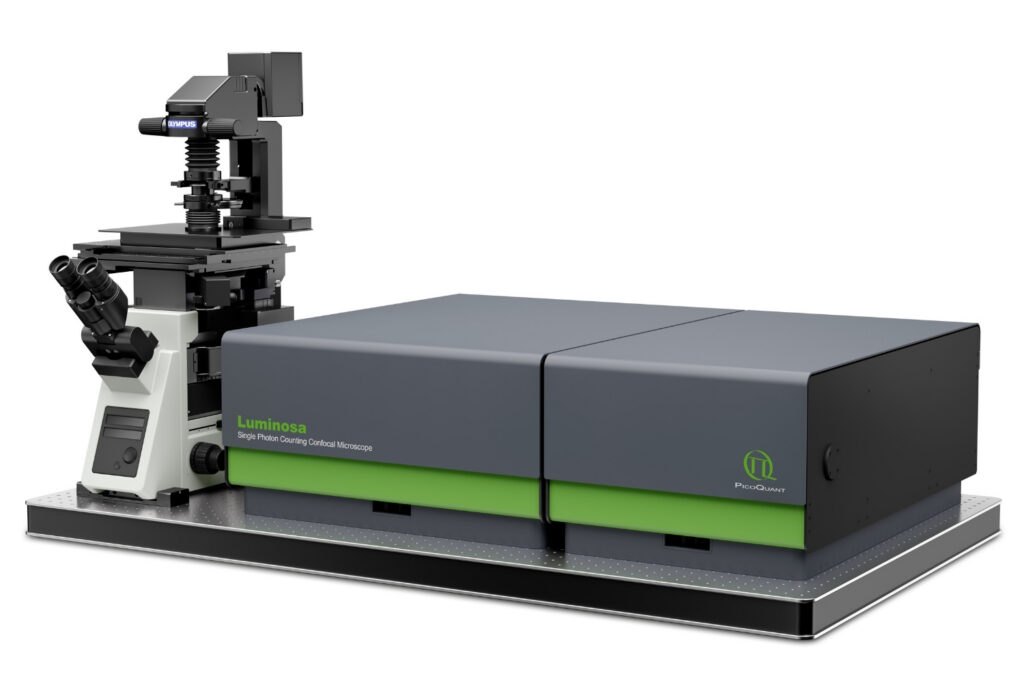

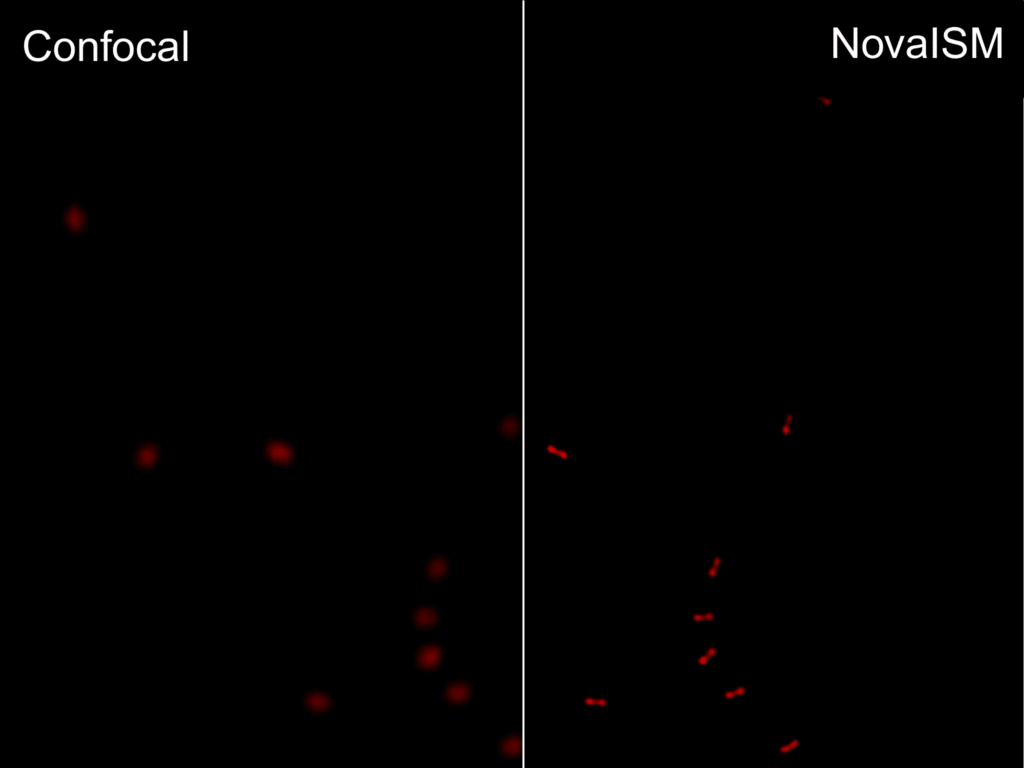
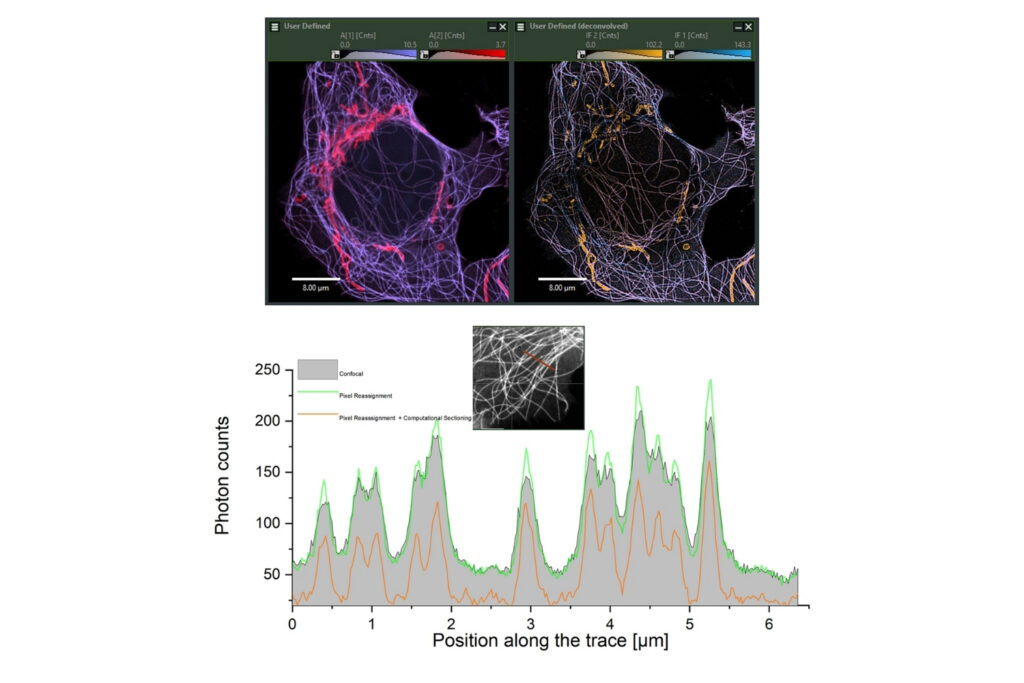
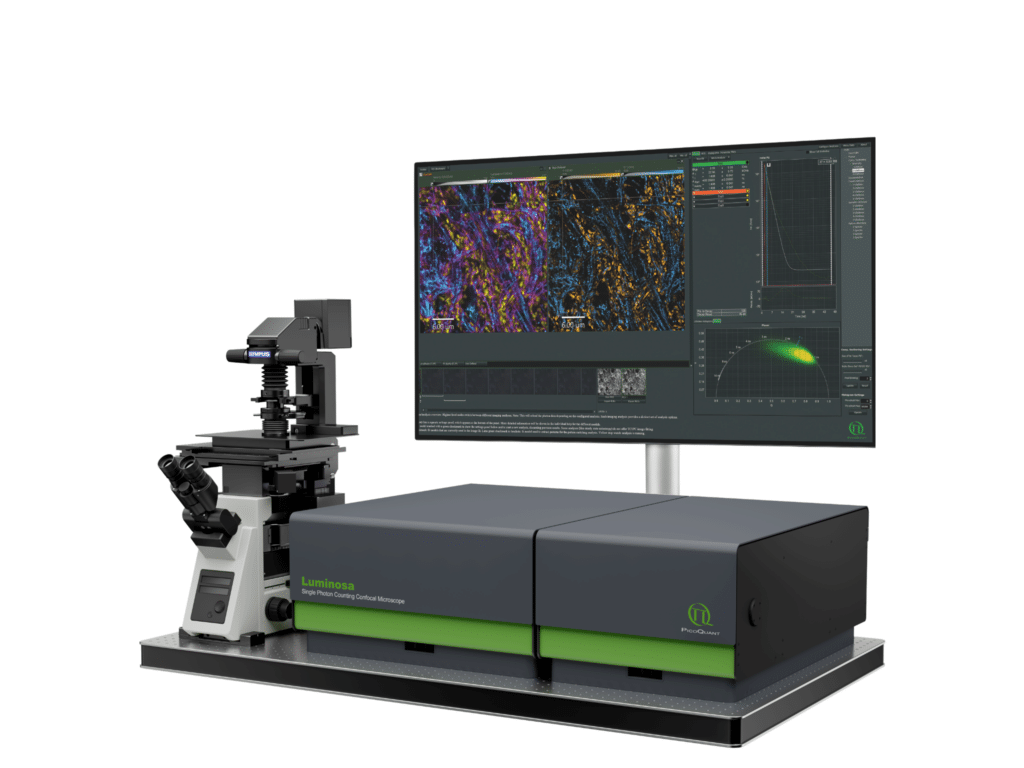
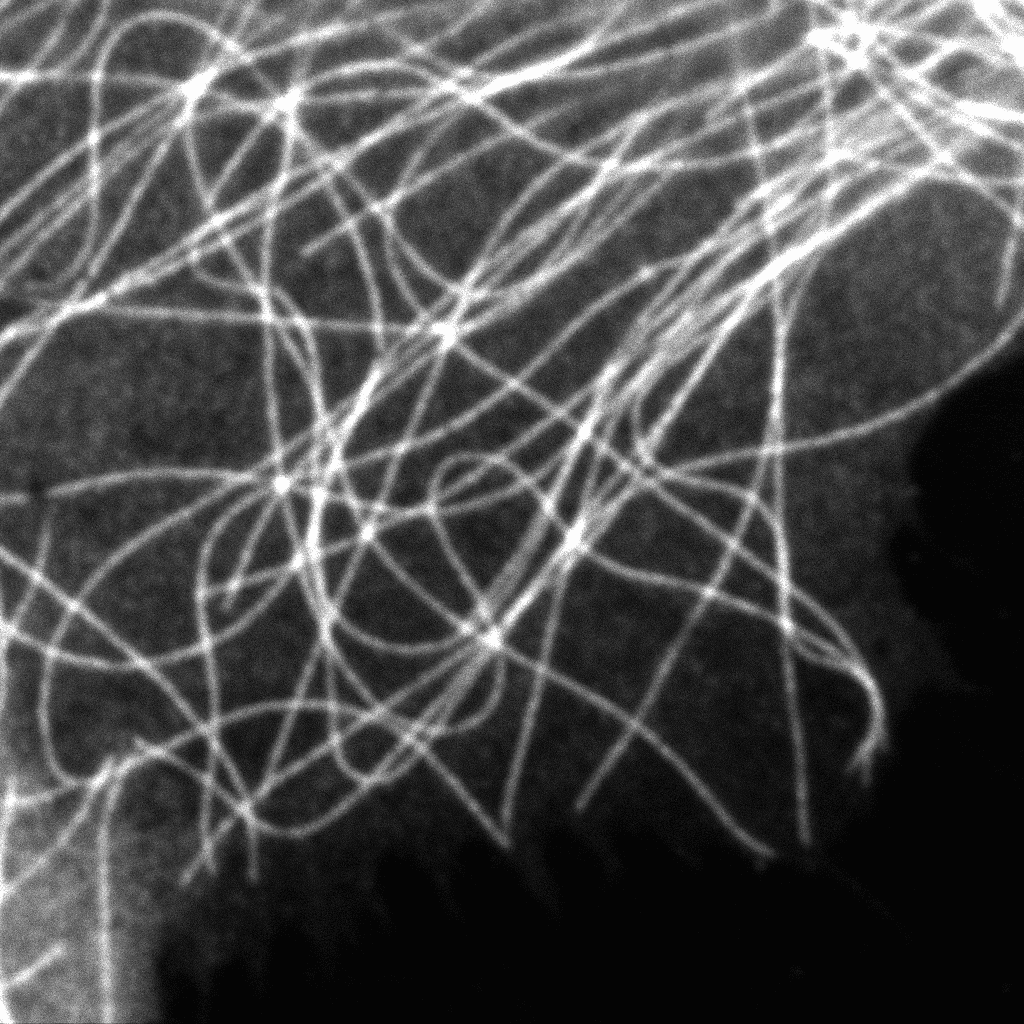
Luminosa
Complete confocal fluorescence microscope that empowers researchers to advance quantitative functional imaging from individual molecules to cells and tissues.
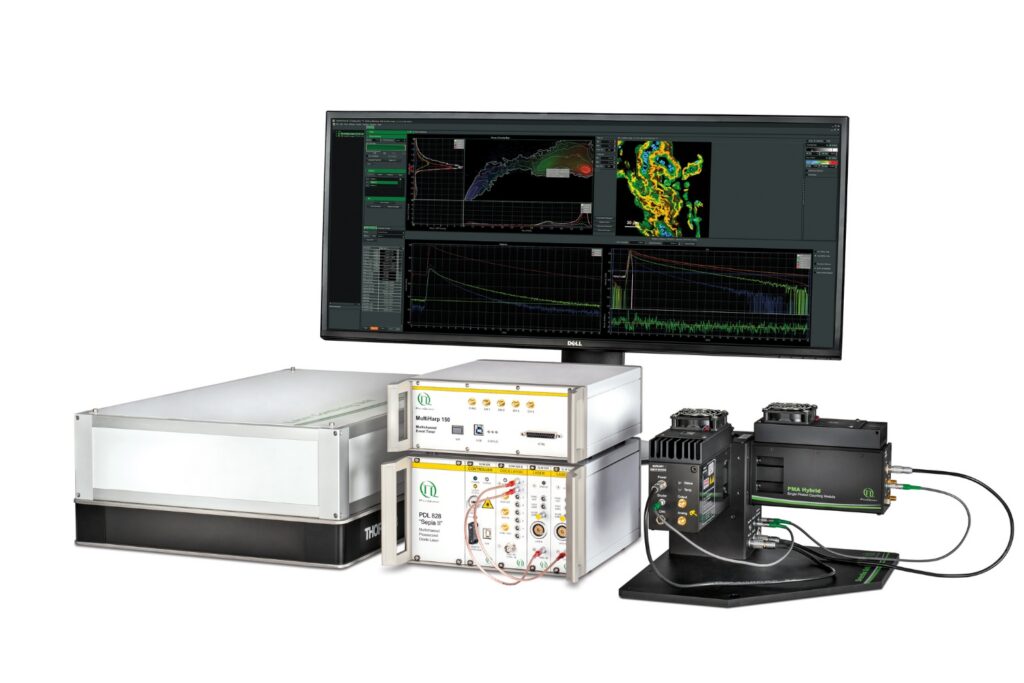

LSM Upgrade Kit
Compact FLIM and FCS upgrade kit that adds advanced functional imaging and correlation analysis to existing laser scanning microscopes.
- All Microscope Systems>220 Z. PicoQuant designs and manufactures spectrometers that range from compact table-top spectrometers for teaching or daily routine work to modular high-end systems with exact timing down to a few picoseconds.
- Spectrometer Systems
Most popular products
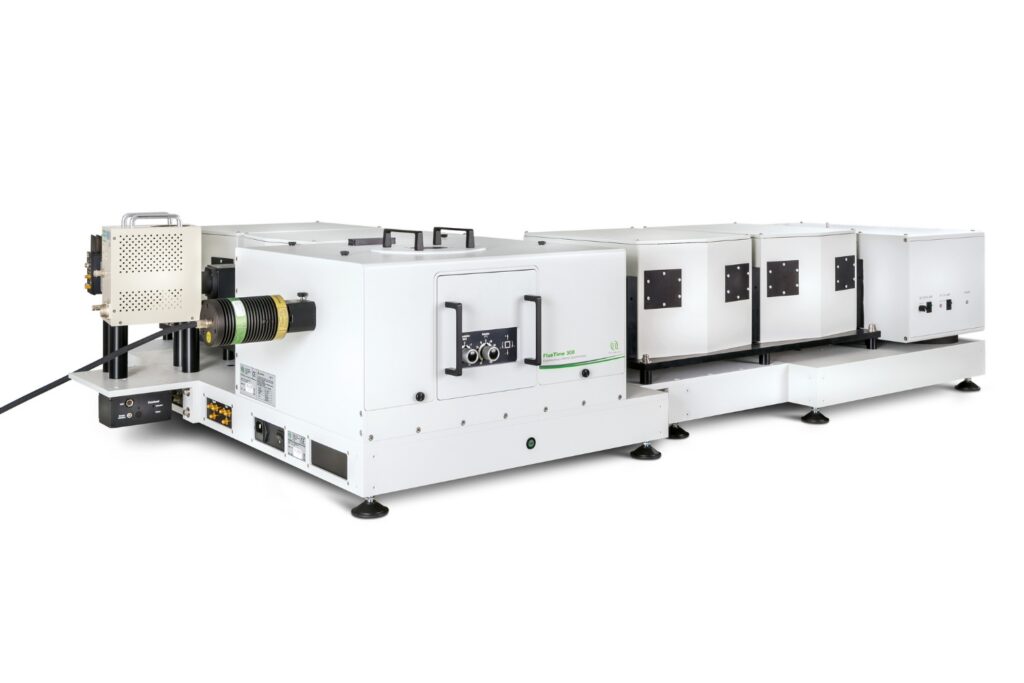

FluoTime 300
Designed for flexible, sensitive, and precise steady-state and time-resolved spectroscopy across the UV to NIR range and time scales from picoseconds to milliseconds.
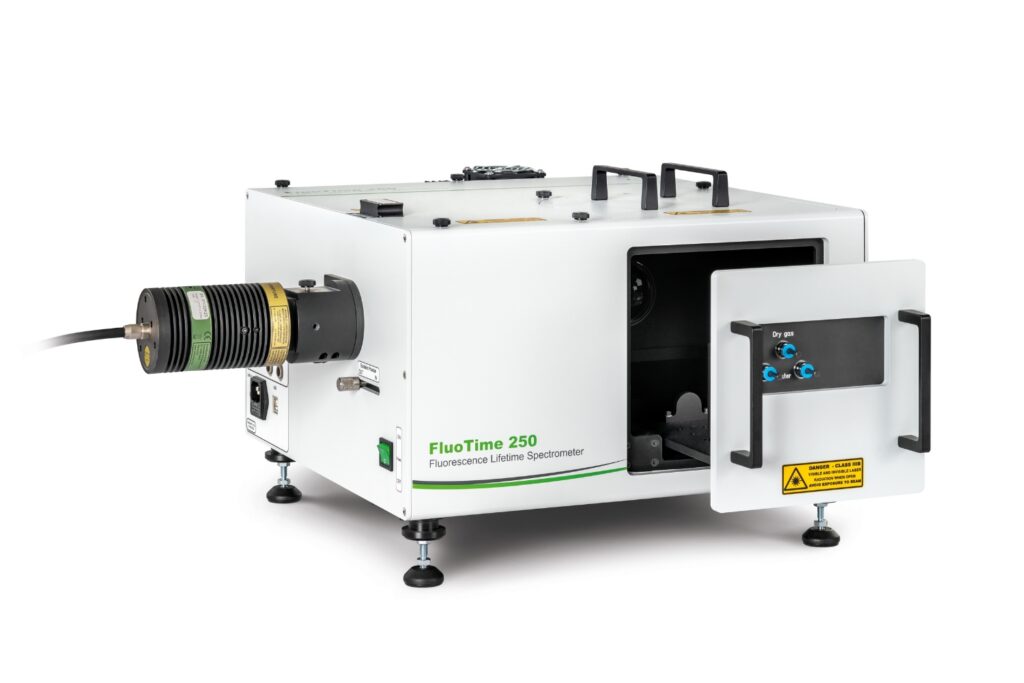

FluoTime 250
Modular lifetime spectrometer designed for flexible fluorescence and photoluminescence measurements in both materials and life science research.

Micro-Photoluminescence Upgrade
Add spectral and time-resolved photoluminescence to your setup through flexible microscope–spectrometer coupling options.
- All Spectrometer Systems>220 Z. PicoQuant designs and manufactures spectrometers that range from compact table-top spectrometers for teaching or daily routine work to modular high-end systems with exact timing down to a few picoseconds.
- Time Tagging and TCSPC Electronics
Most popular products
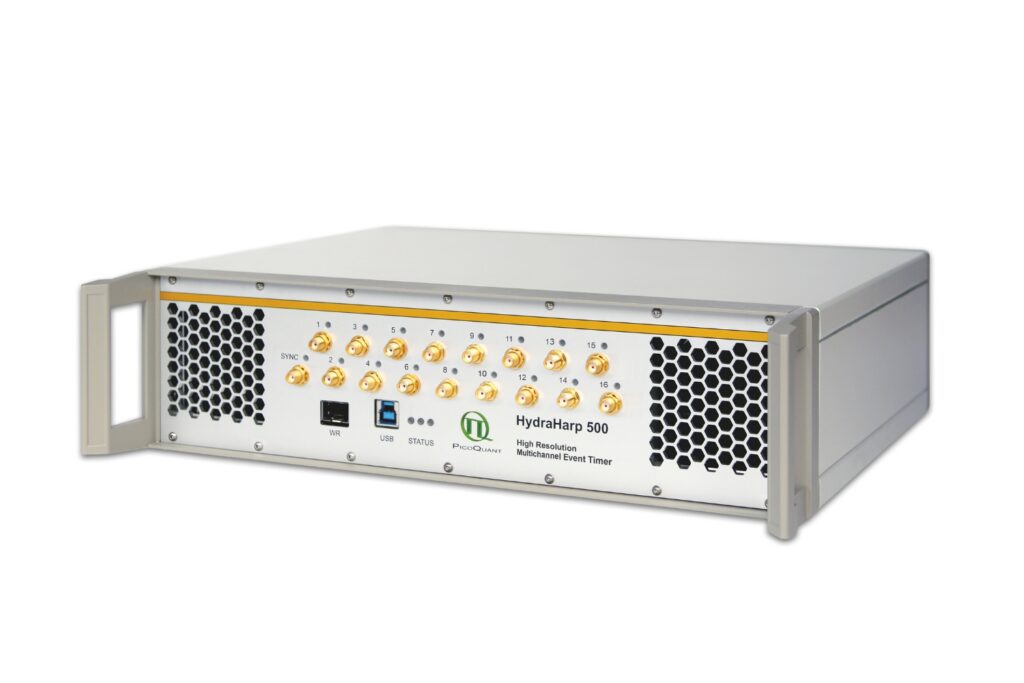

HydraHarp 500
Get the most out of superconducting nanowire detectors in large-scale quantum communication and computing experiments requiring precise multichannel timing.
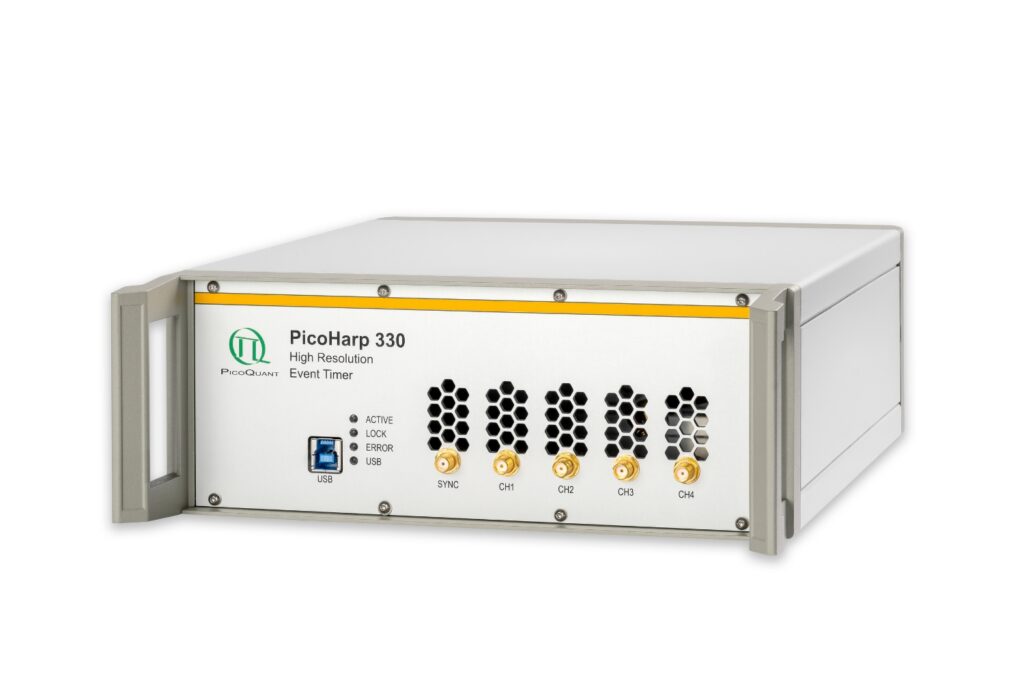

PicoHarp 330
Boost your time-resolved experiments with a flexible, high-precision time tagging and TCSPC unit for materials science and quantum sensing.
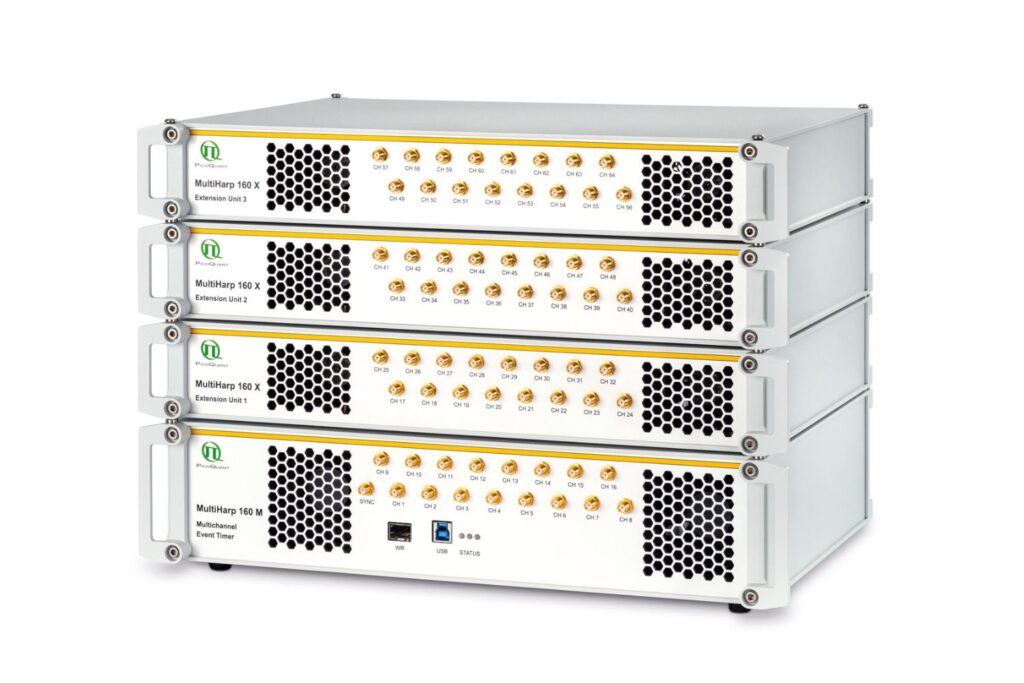

MultiHarp 160
Scale your photonic quantum computing and detector characterization setups while maintaining performance, flexibility, and high data throughput.
- All Time Tagging and TCSPC Electronics>220 Z. PicoQuant designs and manufactures spectrometers that range from compact table-top spectrometers for teaching or daily routine work to modular high-end systems with exact timing down to a few picoseconds.
- Pulsed Lasers, LEDs, and Drivers
Most popular products
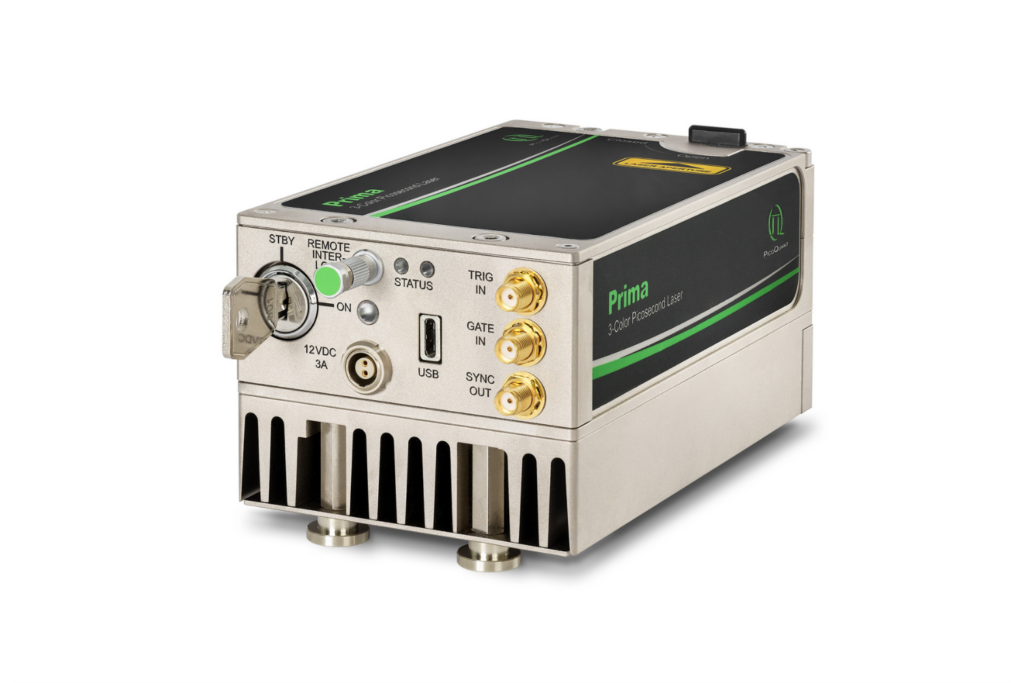

Prima
Compact 3-color picosecond laser delivering flexible ns to ms excitation with cost-effective multicolor performance and straightforward operation.
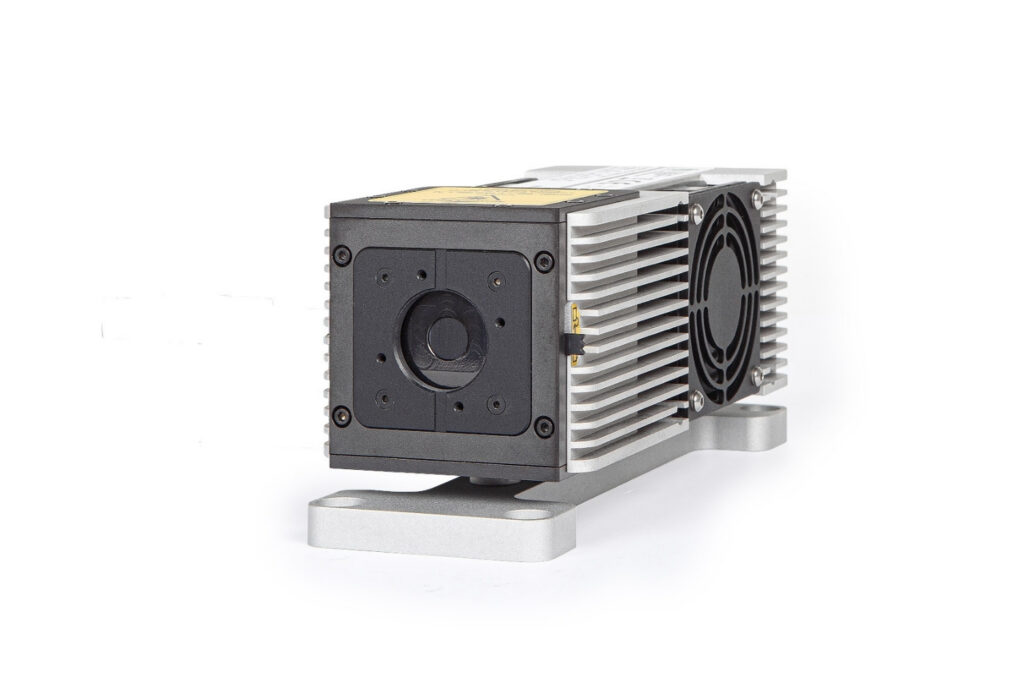

LDH-I Series
Smart picosecond laser diode heads covering UV-A to NIR, providing the right combination of power, pulse width, and diode type for any time-resolved technique.
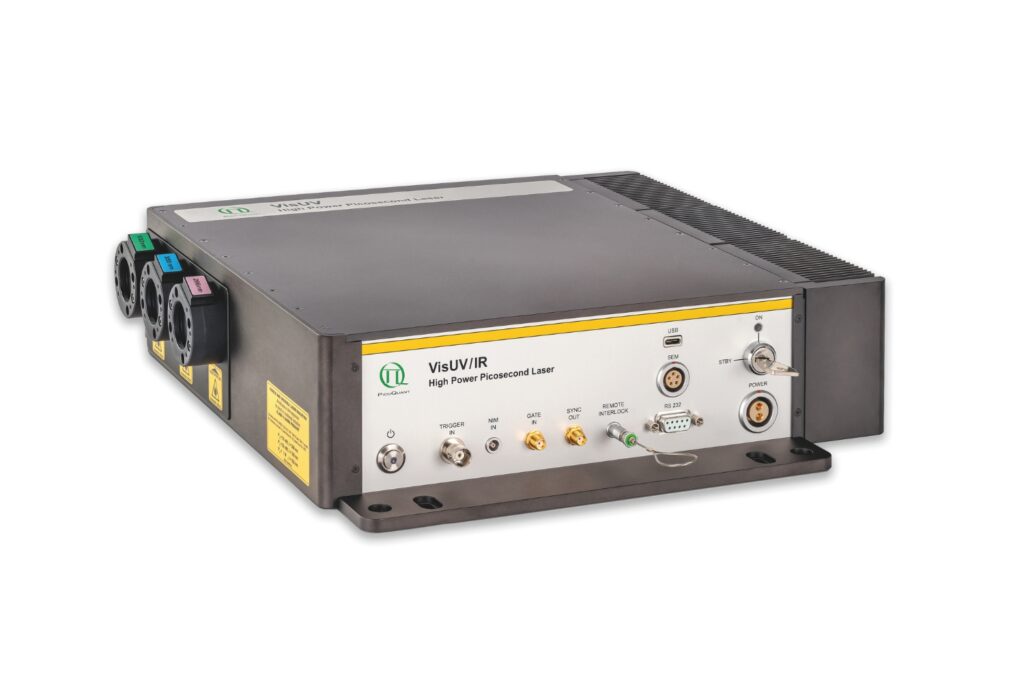

VisUV
VisUV provides clean short pulses and stable timing across key UV and visible wavelengths, including deep UV lines as well as 488 nm and 532 nm.
- All Pulsed Lasers, LEDs, and DriversPulsed lasers and LED sources engineered for picosecond precision, stable output, and flexible use across life science, materials science, quantum optics and metrology.
- Photon Counting Detectors
Most popular products
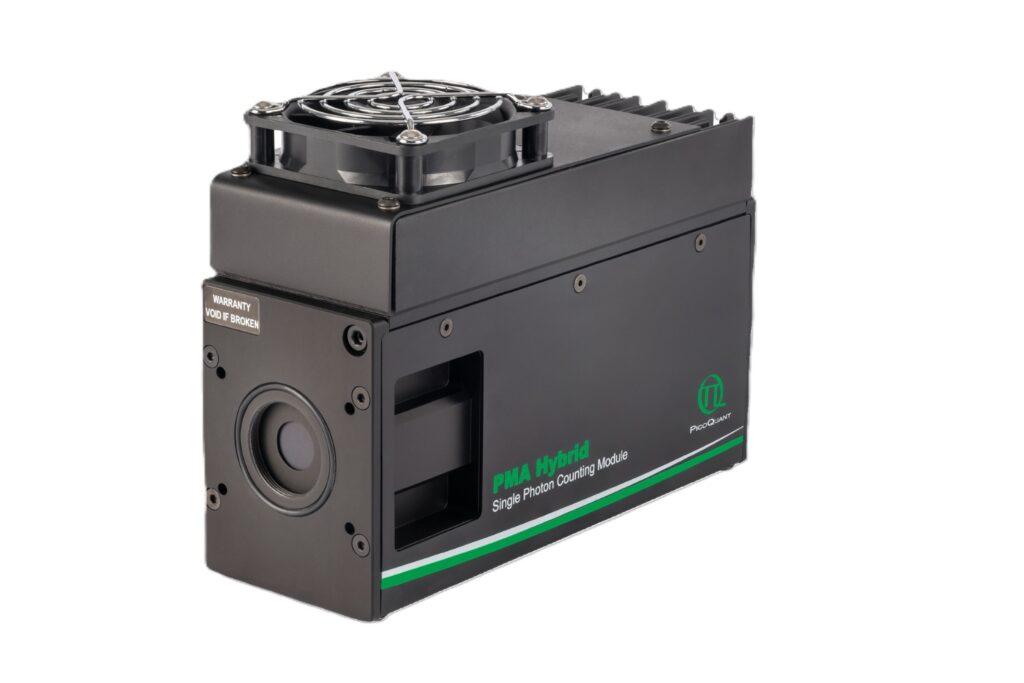

PMA Hybrid Series
Enhance your single-photon counting experiments with wide dynamic range and excellent timing precision in the UV and visible even at the highest count rates.
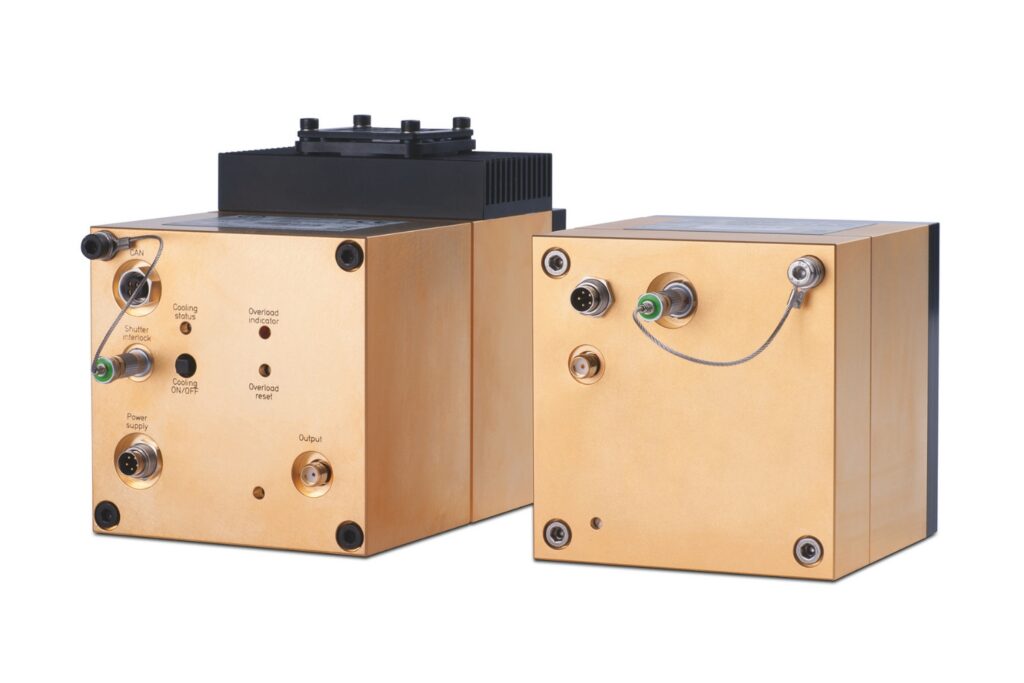

PMA Series
Capture even the weakest signals over large areas with maximum dynamic range and enhanced low-light sensitivity in a compact detector design.
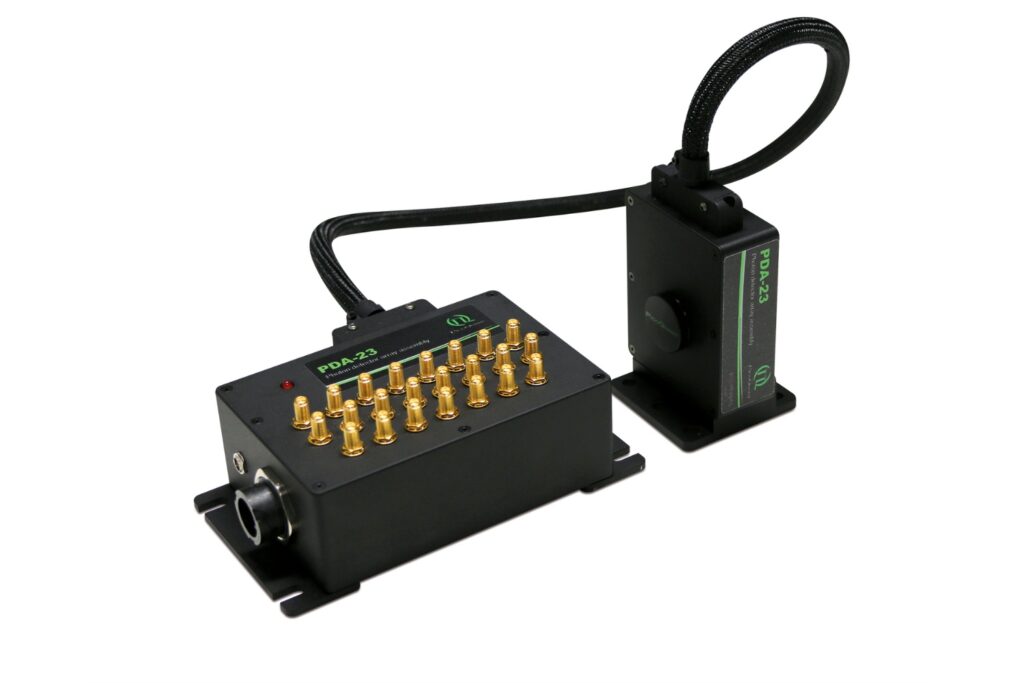

PDA-23
Unlock spatially resolved single-photon detection with a 23-pixel SPAD array, combining low dark counts and precise time tagging for advanced experiments.
- All Photon Counting Detectors>220 Z. PicoQuant designs and manufactures spectrometers that range from compact table-top spectrometers for teaching or daily routine work to modular high-end systems with exact timing down to a few picoseconds.
- Software
Most popular products
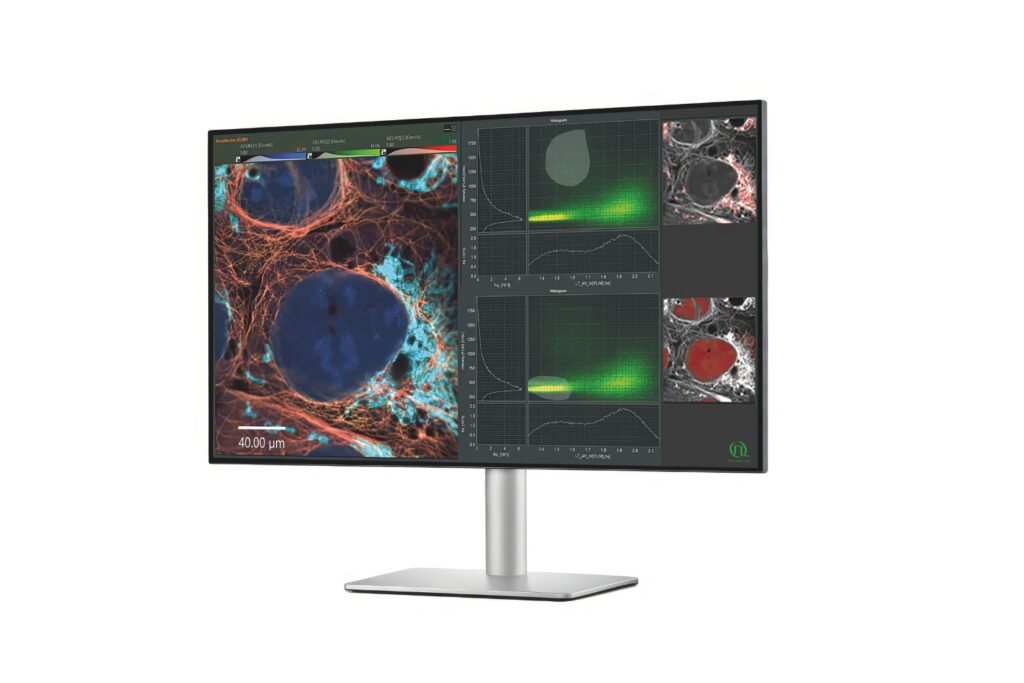
NovaFLIM
Advanced FLIM analysis software for fast, accurate interpretation of lifetime imaging data.
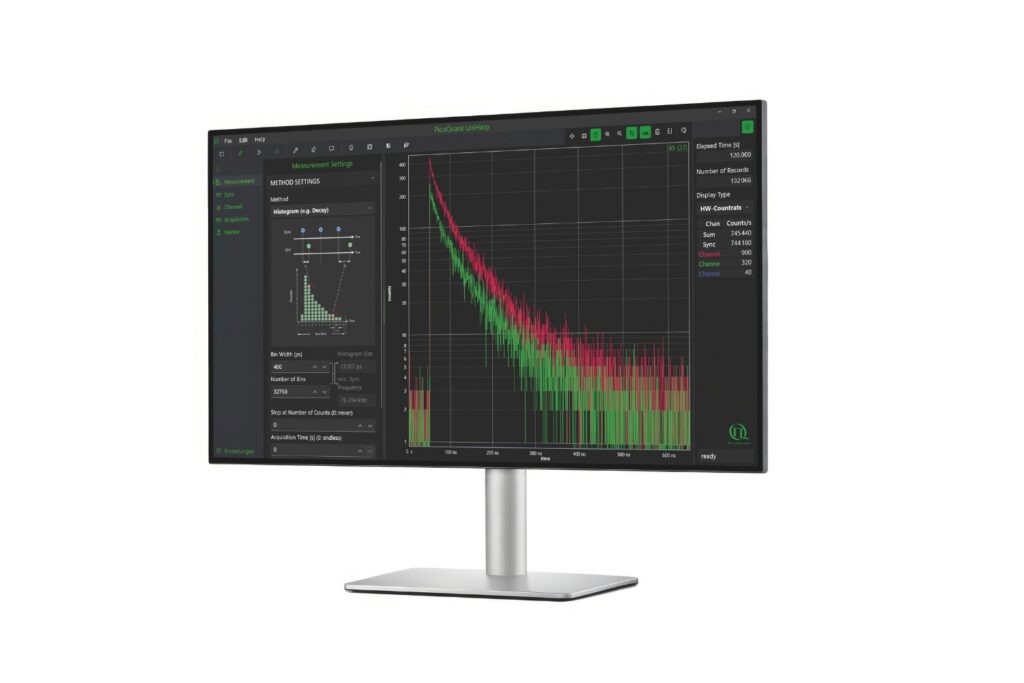
UniHarp
Intuitive, free software solution for real-time, high-precision photon data acquisition, visualization, and initial data analysis.
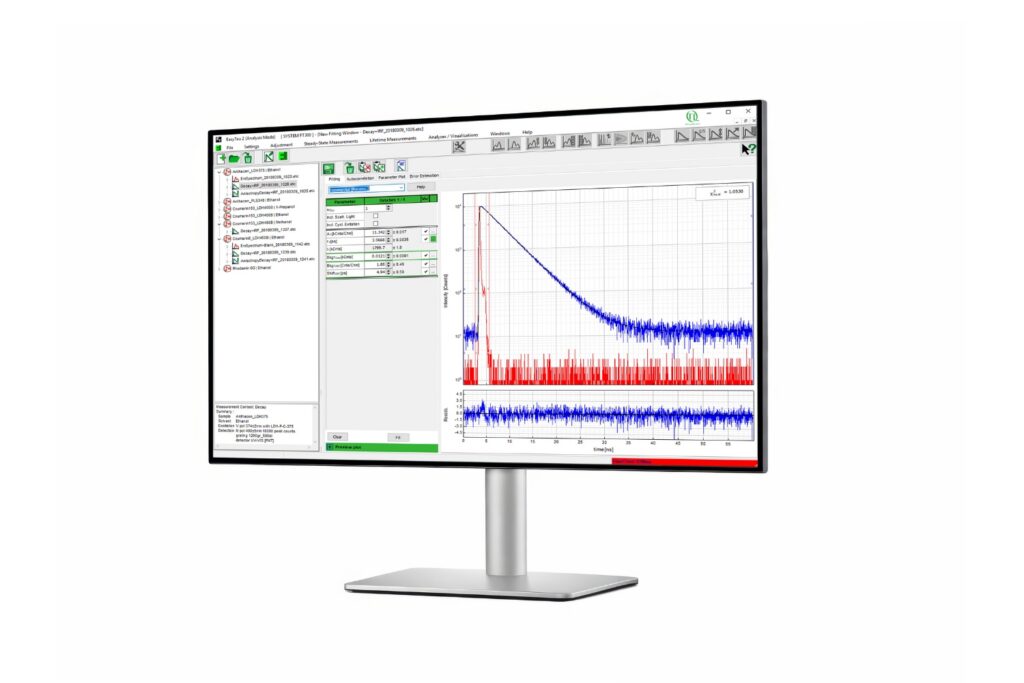
EasyTau 2
Advanced software for time-resolved fluorescence acquisition and analysis.
- All Software>220 Z. PicoQuant designs and manufactures spectrometers that range from compact table-top spectrometers for teaching or daily routine work to modular high-end systems with exact timing down to a few picoseconds.
- Product Overview
Application & Methods
- Life Science
Most popular applications & methods
- All Life Science
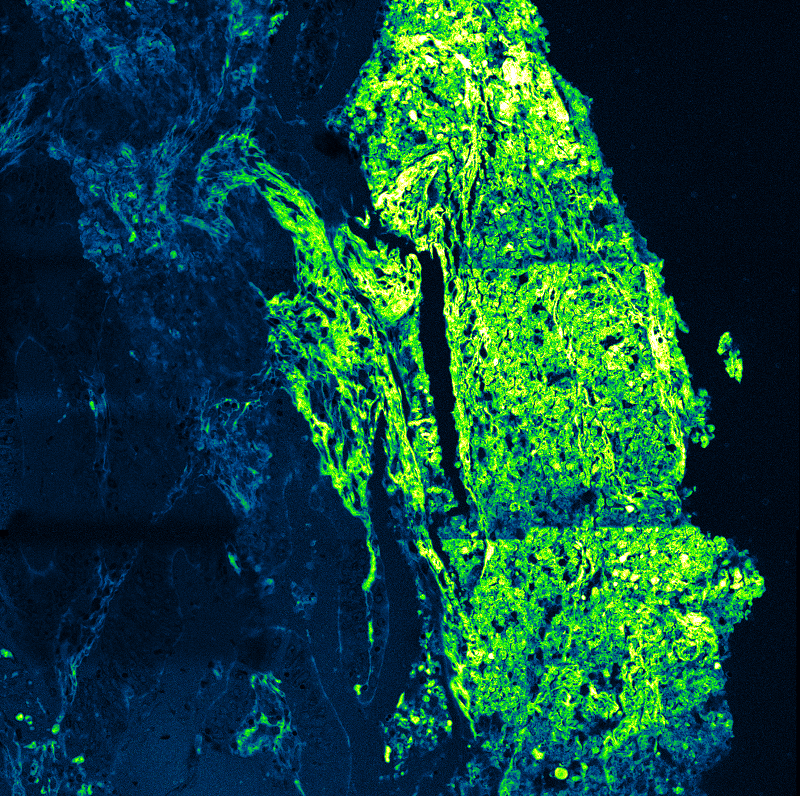
Fluorescence Lifetime Imaging Microscopy (FLIM)
An imaging technique that uses fluorescence lifetimes to generate image contrast.
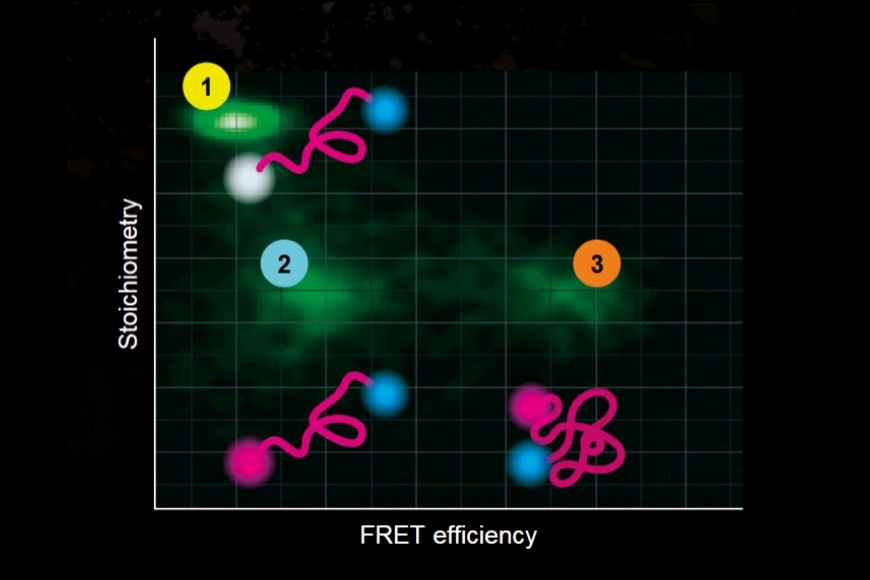
Dynamic Structural Biology
Investigating how proteins dynamically explore multiple conformational states that control biological function.
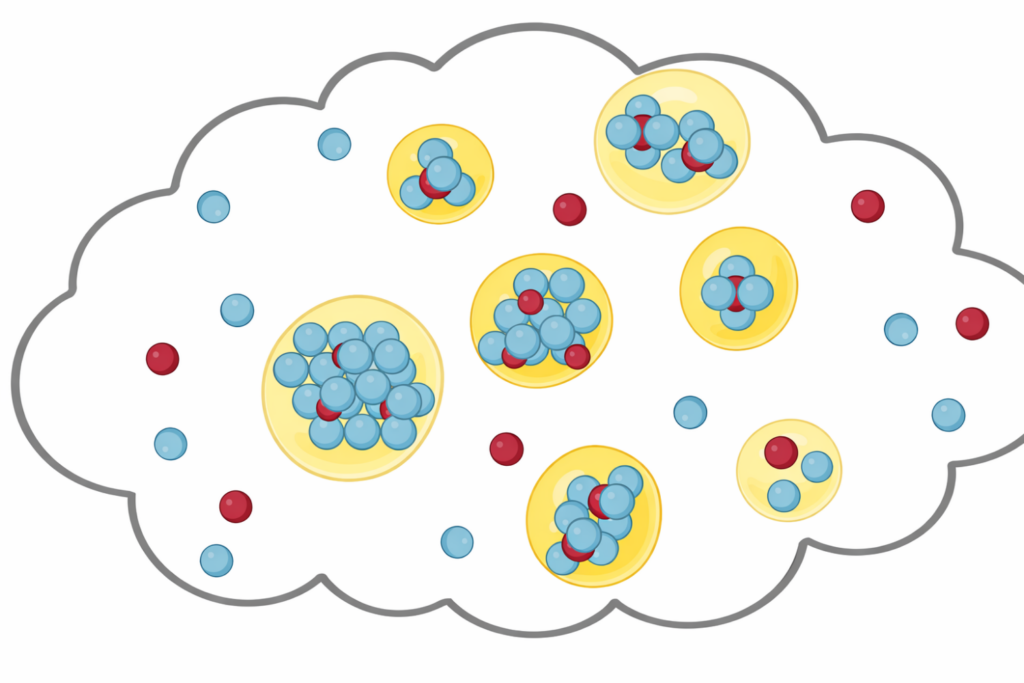
Liquid-Liquid Phase Separation (LLPS)
Investigating how biomolecules separate into dynamic liquid phases to organize cellular space and regulate biological function.
- Materials Science
Most popular applications & methods
- All Materials Science
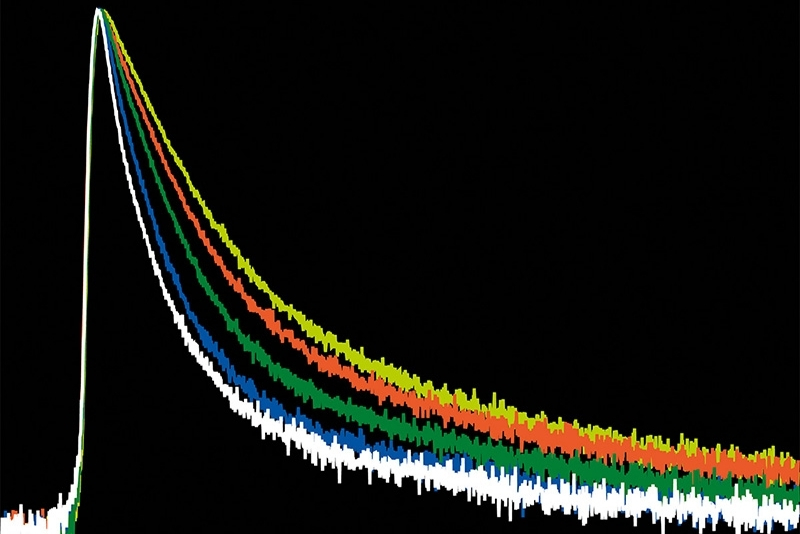
Time-Resolved Photoluminescence (TRPL)
A time-resolved technique that measures photoluminescence lifetimes to reveal excited-state dynamics in materials.
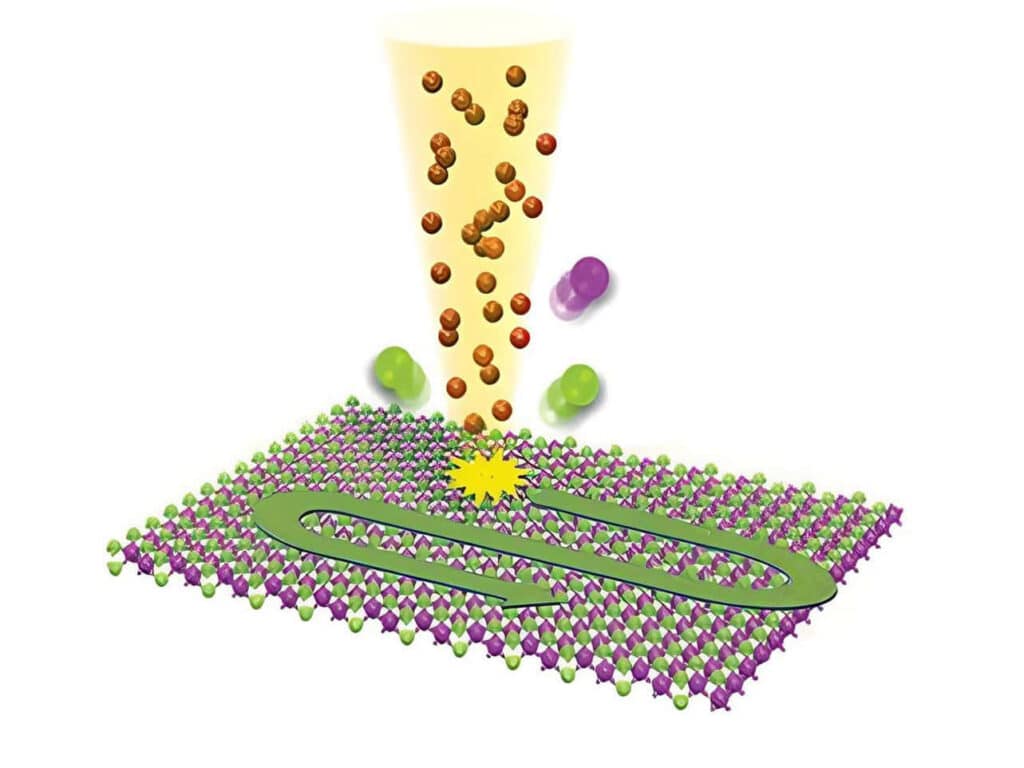
2D Materials Research
Studying exciton dynamics, charge carrier processes, and structural properties through optical and time-resolved characterization methods.

Solar Cell Characterization
Investigating charge-carrier lifetimes and recombination dynamics to enable precise optical characterization of material quality and device performance.
- Quantum Optics
Most popular applications & methods
- All Quantum Optics
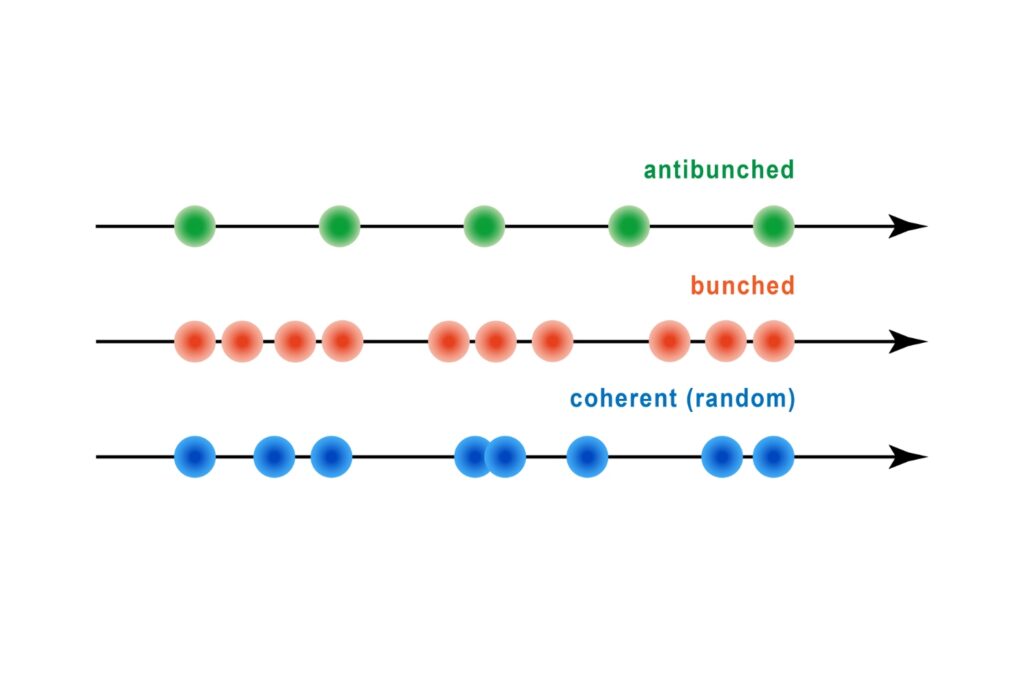
Antibunching
A quantum optical signature revealed by time-resolved photon correlation analysis to identify single-photon emission in materials and nanostructures.
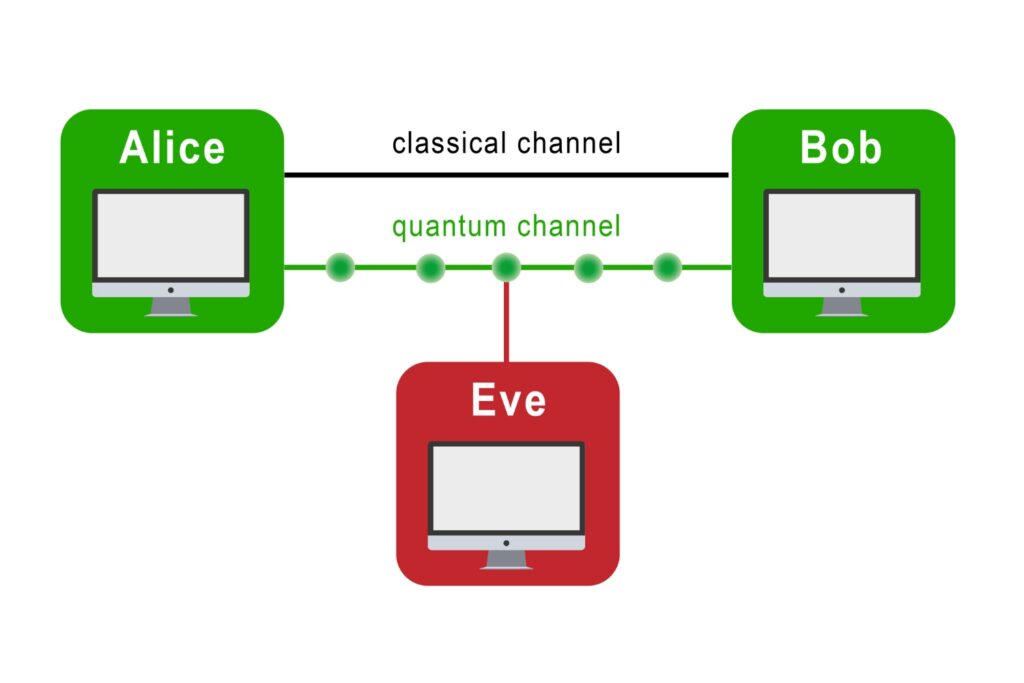
Quantum Communication
The transmission of information using individual photons, using quantum effects to ensure absolute security.
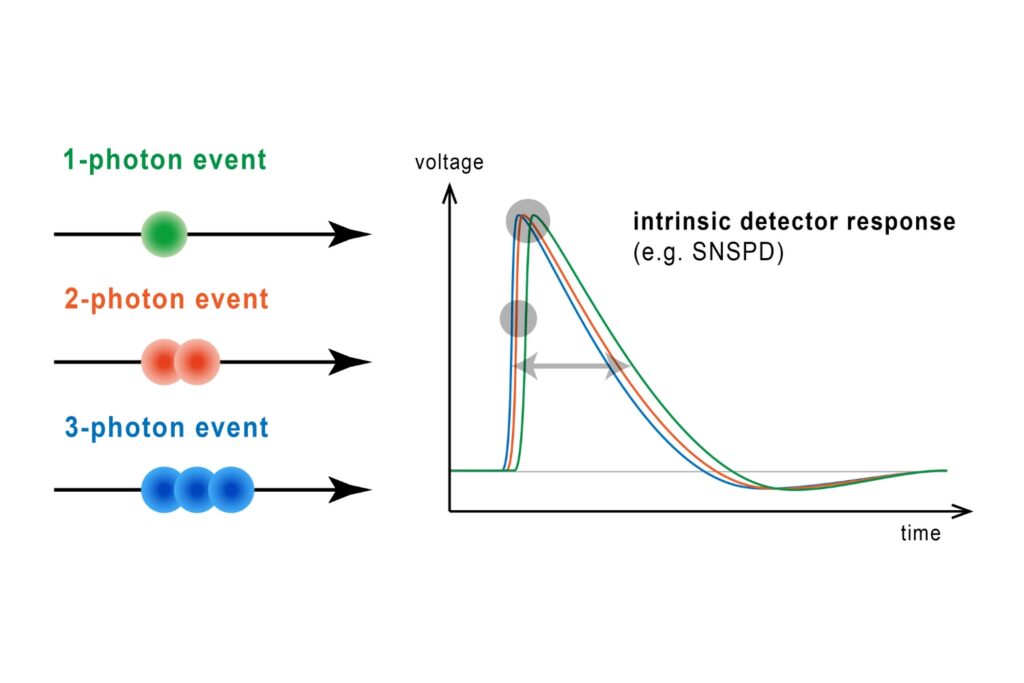
Photon Number Resolution (PNR)
Quantifying photons per detection event enables direct access to photon-number statistics, providing insight into quantum and statistical properties of light.
- Metrology
Most popular applications & methods
- All Metrology
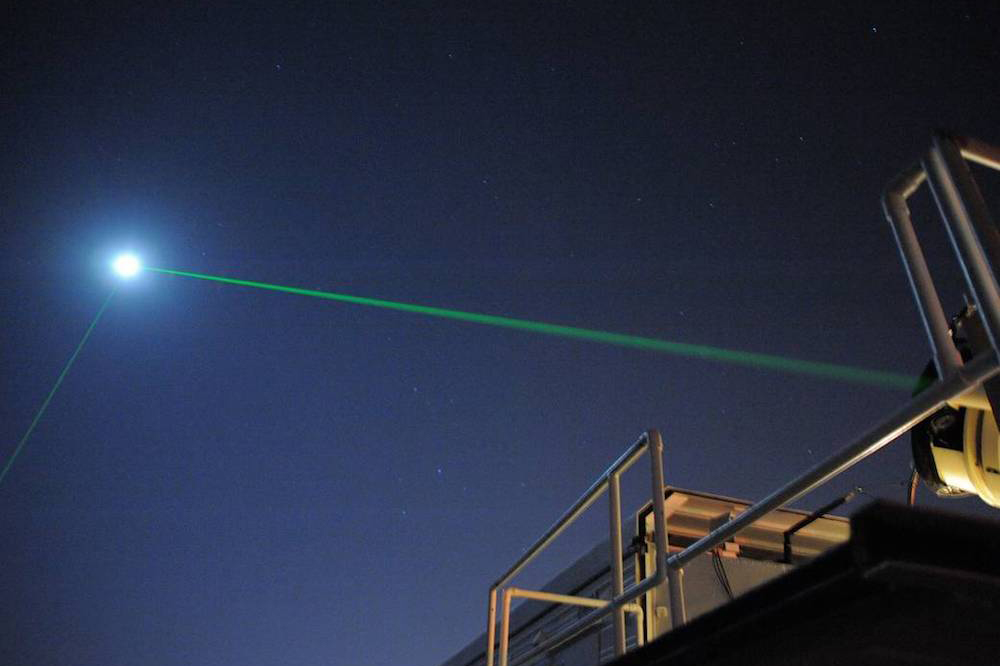

LiDAR, Ranging, and Satellite Laser Ranging (SLR)
An optical technique that analyzes light emission under electrical excitation to reveal electronic properties of electroluminescent materials.
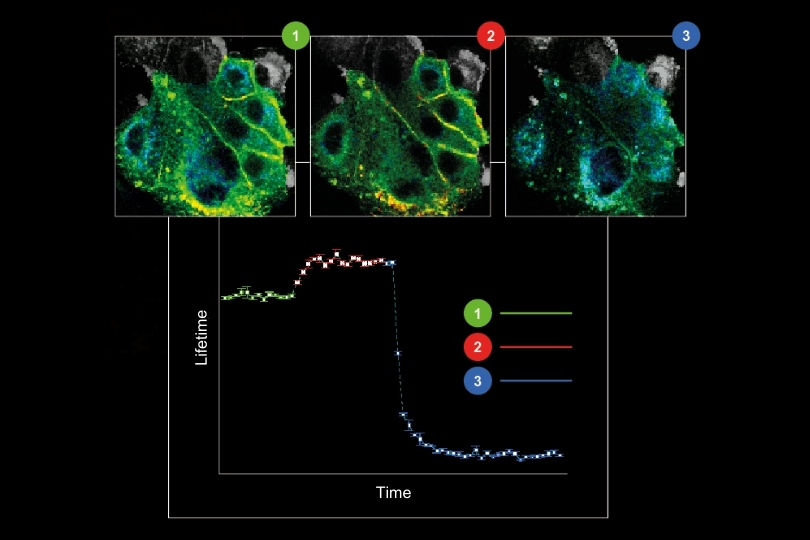
Environmental Sensing
Monitoring environmental signals and trace compounds to understand dynamic changes in natural and engineered environments.
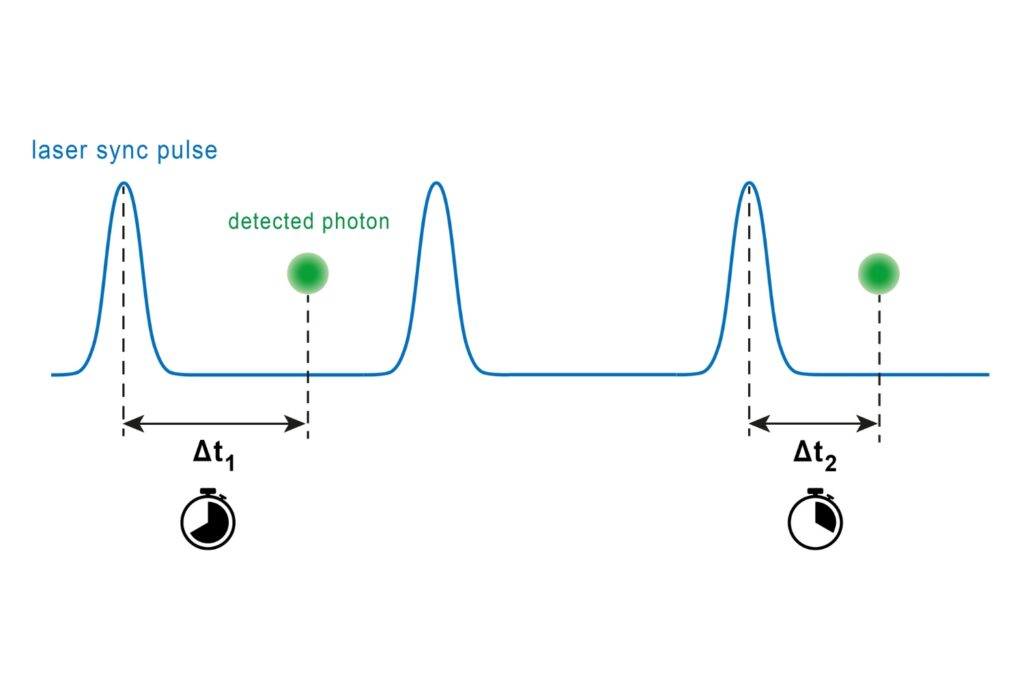
Time-Correlated Single Photon Counting (TCSPC)
A photon timing technique that measures single-photon arrival times to resolve ultrafast dynamics in fluorescence, materials research, and quantum optics.
- All Applications & Methods