Key Highlights
- Quantitative FLIM-FRET enables concentration-independent detection of PRG5 multimerization within confined membrane domains.
- TCSPC-based lifetime imaging resolves nanosecond-scale interaction differences across subcellular regions, including filopodia and neuronal spine-like protrusions.
- Region-specific fluorescence lifetime analysis links biochemical evidence of multimerization to spatially resolved protein organization in living cells.
Spine Formation and the Need for Quantitative Interaction Mapping
Dendritic spines form the structural basis of synaptic plasticity. Their morphology and density are tightly regulated, and abnormalities in spine formation are associated with numerous neurological disorders. Understanding how membrane-associated proteins organize within these highly confined structures is therefore not merely a cell biological question but a clinically relevant one.
Plasticity-related gene 5 (PRG5) is a neuron-specific membrane protein implicated in filopodia formation and the development of spine-like protrusions. Previous work suggested that PRG5 may form dimers or higher-order multimers, yet conventional biochemical assays could not resolve where these complexes assemble inside intact cells. The central challenge was clear: how can protein multimerization be detected and spatially mapped within micrometer-scale membrane domains in living systems.
Why Quantitative FLIM-FRET Was Critical
Protein–protein interactions are often studied using intensity-based FRET approaches. However, intensity measurements depend strongly on fluorophore concentration, expression levels, spectral bleed-through, and local excitation conditions. These factors become particularly problematic when investigating spatially restricted interactions at the plasma membrane or within thin protrusions.
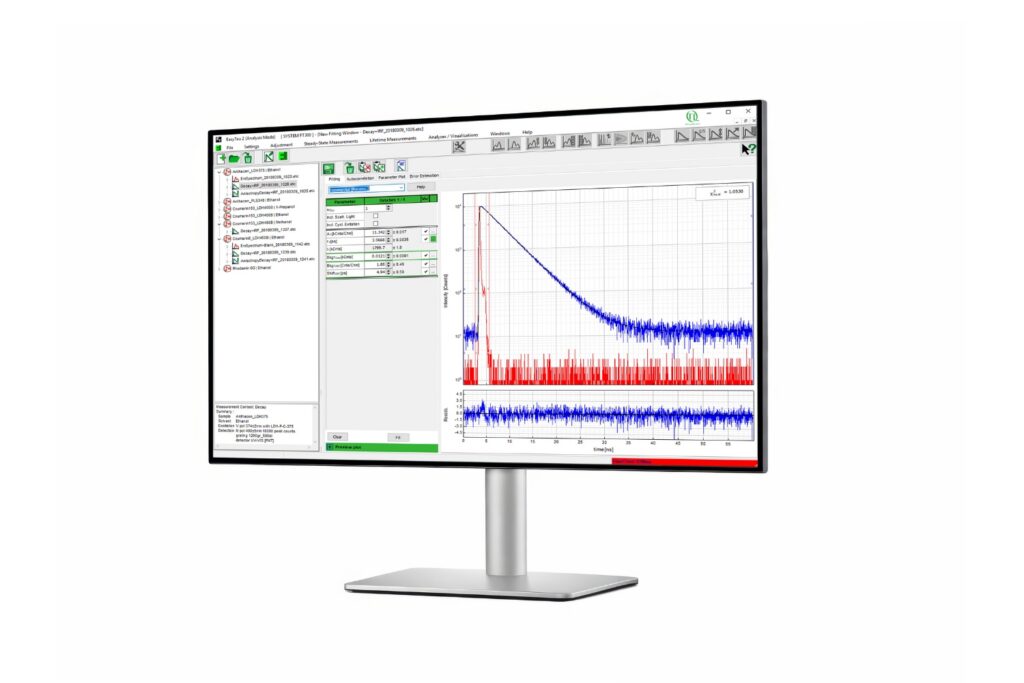
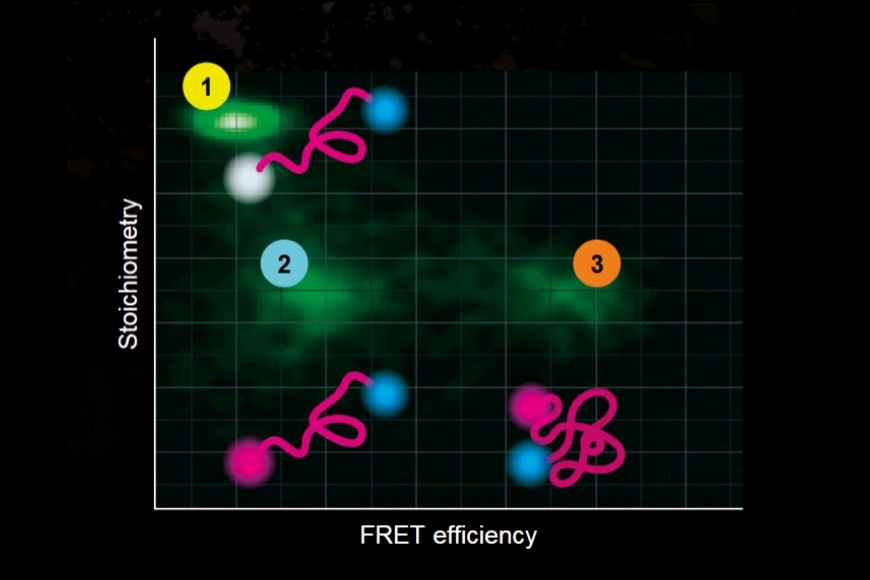
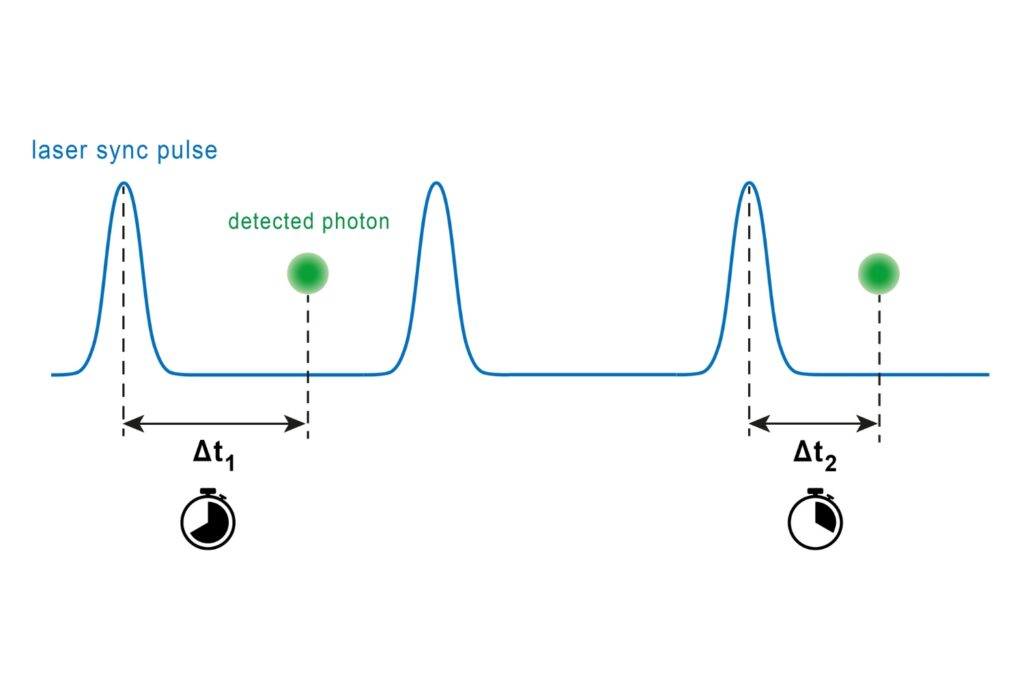
In this study, Köper et al., Front. Mol. Biosci. (2024), combined fluorescence lifetime imaging microscopy (FLIM) with Förster resonance energy transfer (FRET) to overcome these limitations. Rather than relying on intensity ratios, FLIM-FRET quantifies the reduction of the donor fluorescence lifetime that occurs when an acceptor fluorophore is brought into close proximity. Because FRET requires intermolecular distances below approximately 10 nanometers, lifetime shortening provides a direct, concentration-independent proxy for molecular interaction.
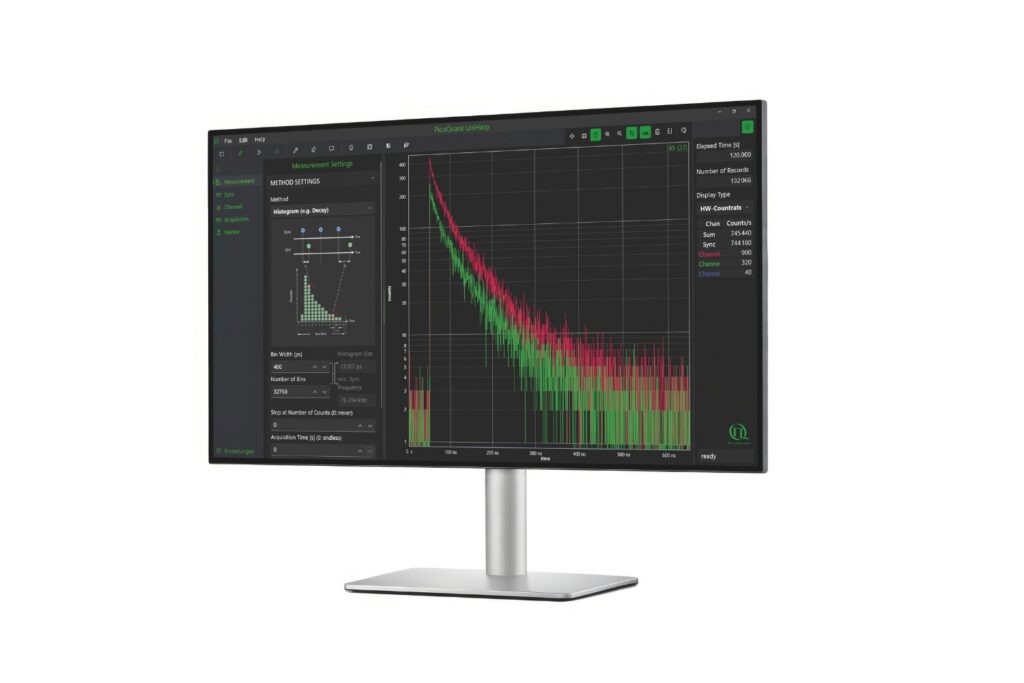
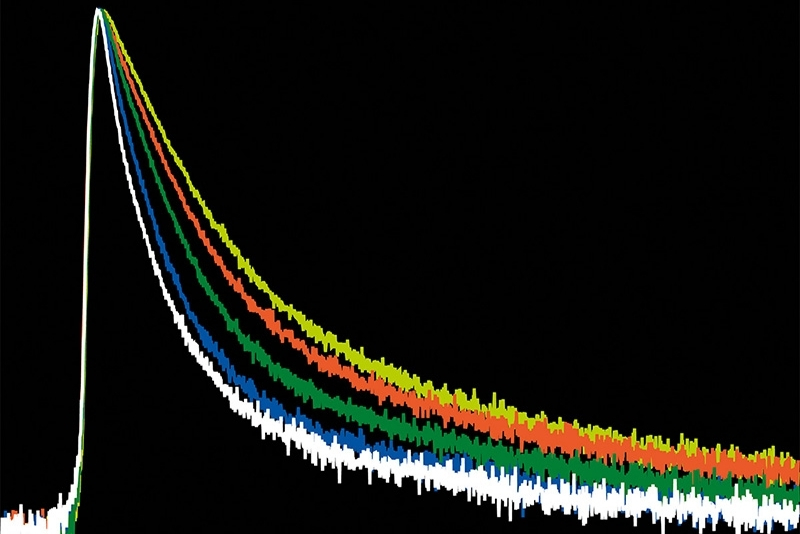
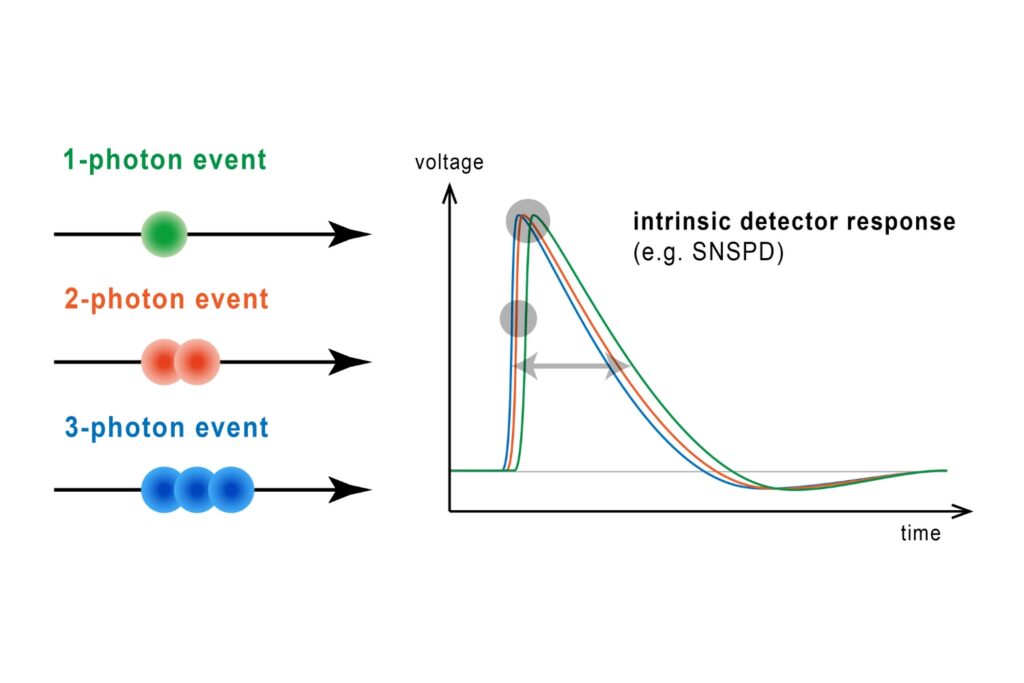
The system operated in time-correlated single photon counting mode with a temporal resolution of 10 picoseconds. Photon counting enables robust decay statistics, while multi-exponential amplitude-weighted lifetime fitting allows reliable discrimination of nanosecond-scale lifetime shifts. In the context of membrane-confined protein assemblies, this quantitative precision was essential.
Core Findings of the Study
Biochemical analyses first established that PRG5 is present not only as a monomer but also in dimeric and higher molecular weight forms. Solubility and deglycosylation assays demonstrated that these species were not insoluble aggregates but soluble, N-glycosylated forms. Co-immunoprecipitation further supported the existence of PRG5 homo-dimers. These experiments indicated multimerization but provided no spatial resolution.
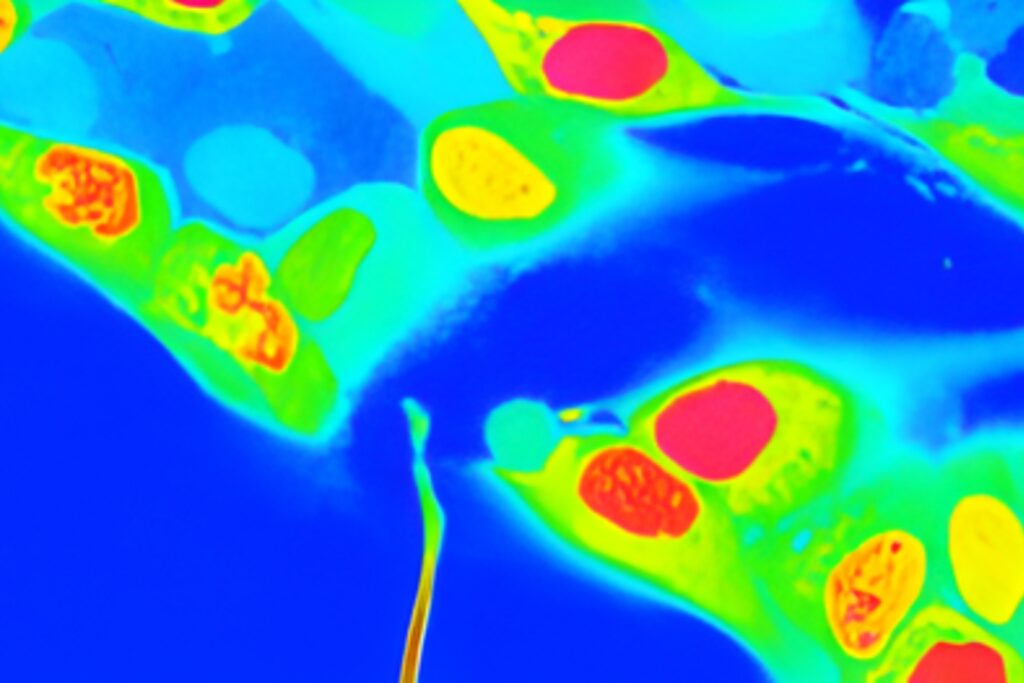
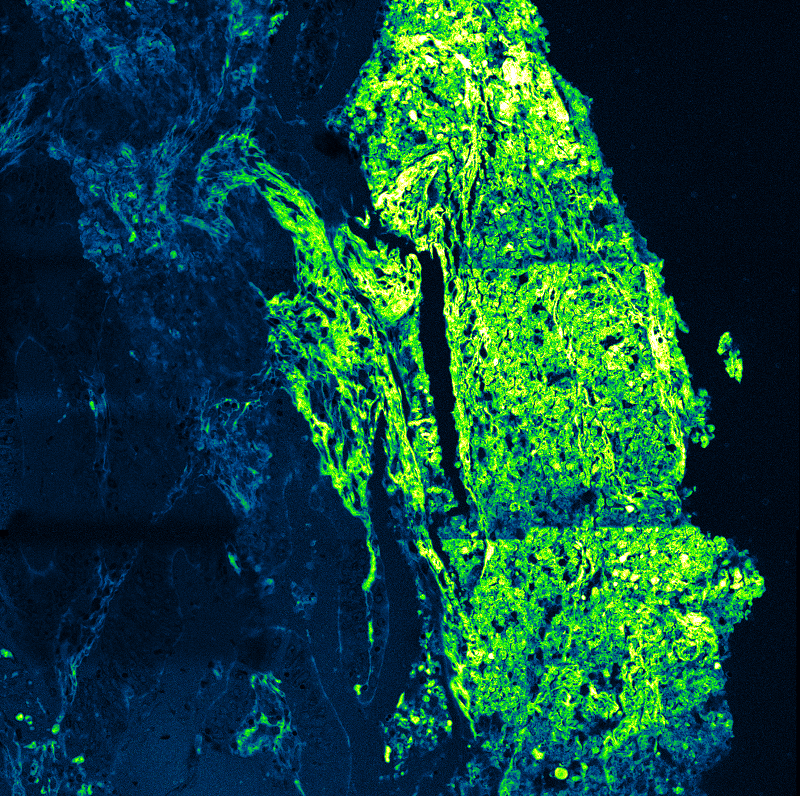
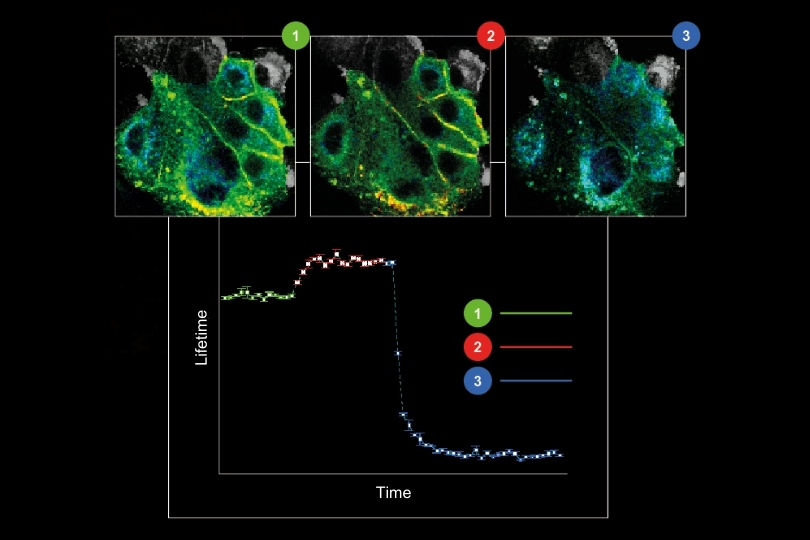
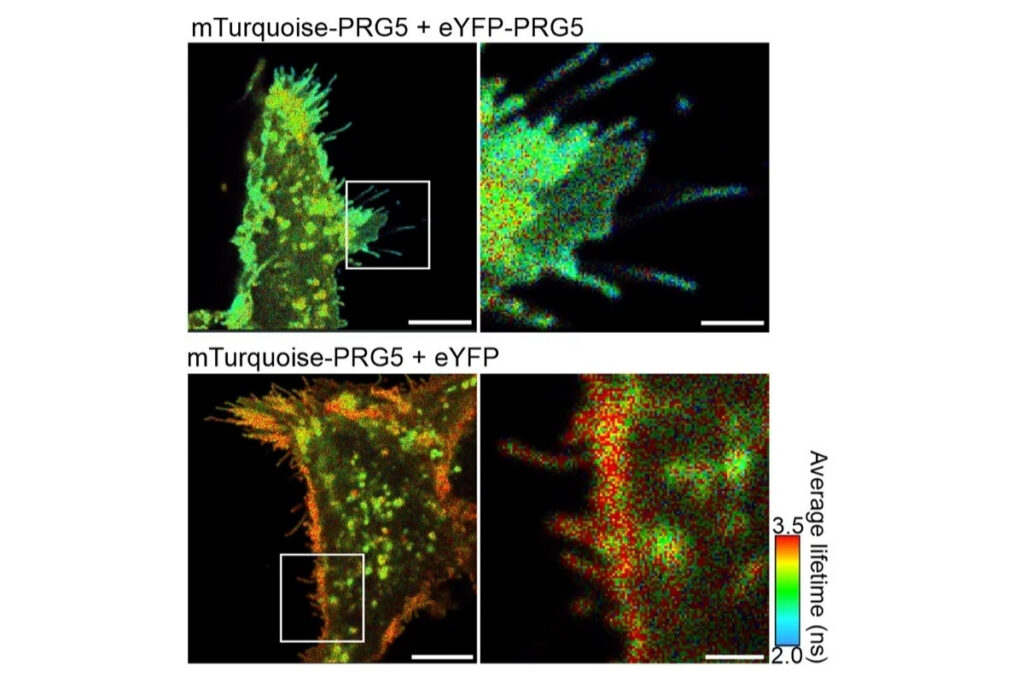
Live-cell FLIM-FRET measurements in HEK293T cells revealed a pronounced reduction in donor lifetime when PRG5 donor and acceptor constructs were co-expressed, corresponding to FRET efficiencies of approximately 50 percent for C-terminally tagged constructs. Lifetime shortening was localized specifically at the plasma membrane, both in regions with and without filopodia. When filopodia were analyzed in greater detail, shorter lifetimes were observed within these protrusions, with a tendency toward further reduction at distal regions.
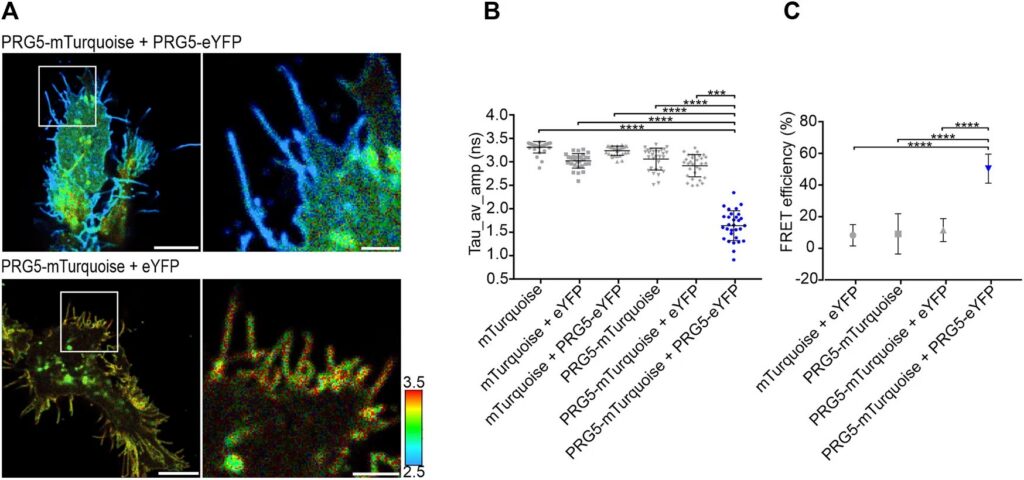
The spatial specificity became even more pronounced in primary hippocampal neurons. Overexpression of PRG5 induced spine-like membrane structures, consistent with earlier findings. FLIM-FRET analysis demonstrated significant donor lifetime shortening within these protrusions, with the strongest reduction at the distal tips. This region-resolved analysis revealed that PRG5 multimers are not uniformly distributed but are enriched in defined subcellular compartments associated with membrane protrusion dynamics.
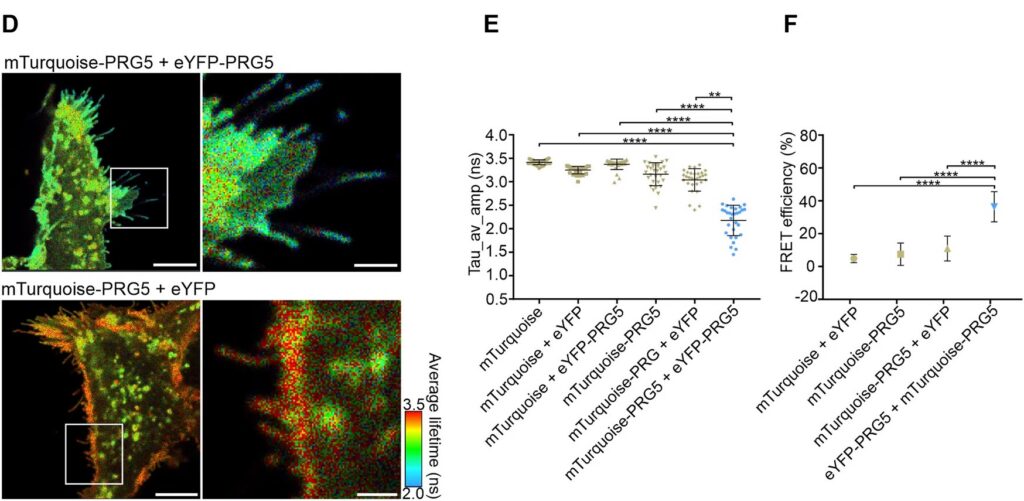
Together, these results demonstrate that quantitative lifetime imaging enables the detection and spatial mapping of protein multimerization within highly confined cellular microstructures.
Why TCSPC-Based Lifetime Imaging Enabled This Analysis
The ability to detect approximately one-nanosecond lifetime shifts in restricted membrane domains depends on both temporal resolution and statistical robustness. Time-correlated single photon counting provides precise photon timing and accurate reconstruction of fluorescence decay curves. In this study, a 10 picosecond timing resolution ensured that subtle lifetime differences could be reliably quantified.
Because FLIM-FRET is independent of fluorophore concentration, it allows comparison between different membrane regions without intensity normalization artifacts. This concentration independence was critical for distinguishing true interaction-driven lifetime changes from variations in local expression or membrane topology. The combination of TCSPC electronics and confocal microscopy thus enabled a level of quantitative interaction mapping that would be difficult to achieve with intensity-based approaches.
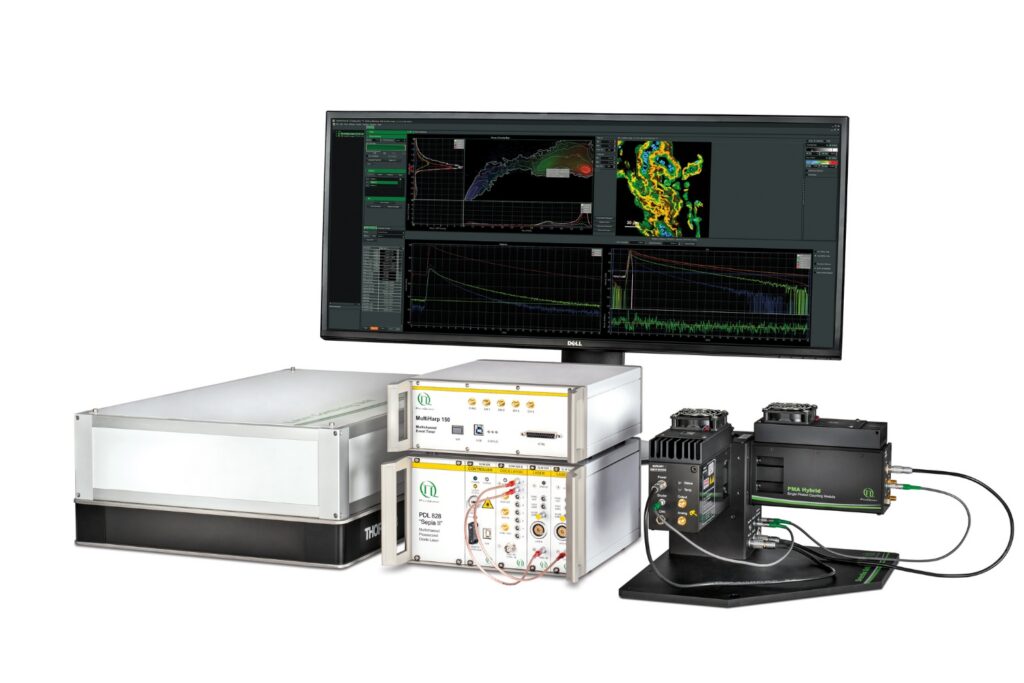
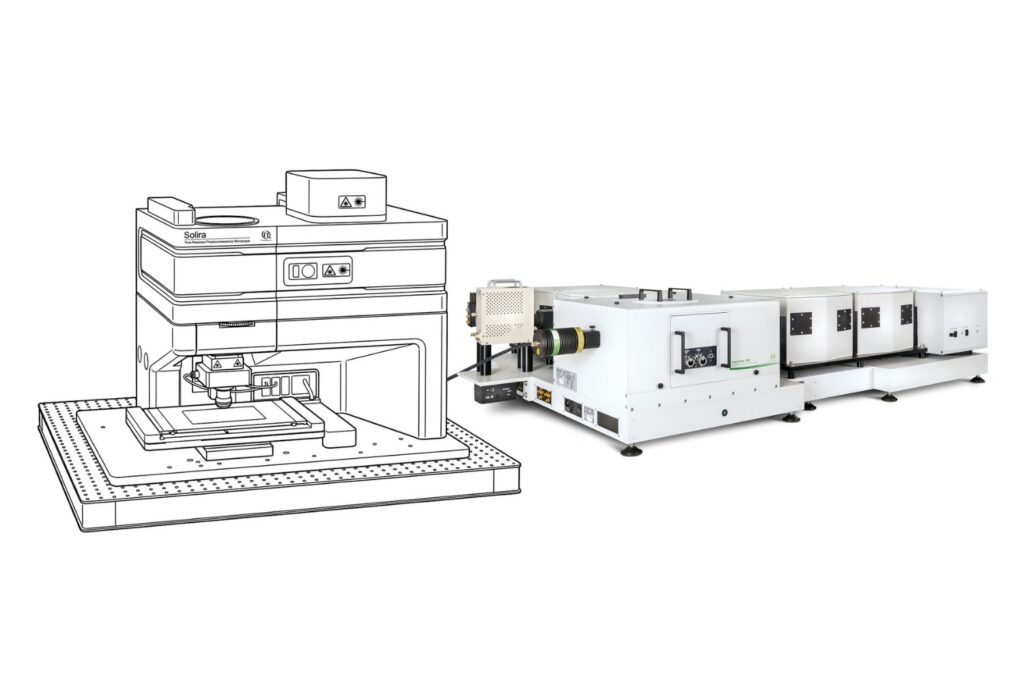
Instrumentation Used in This Study by PicoQuant
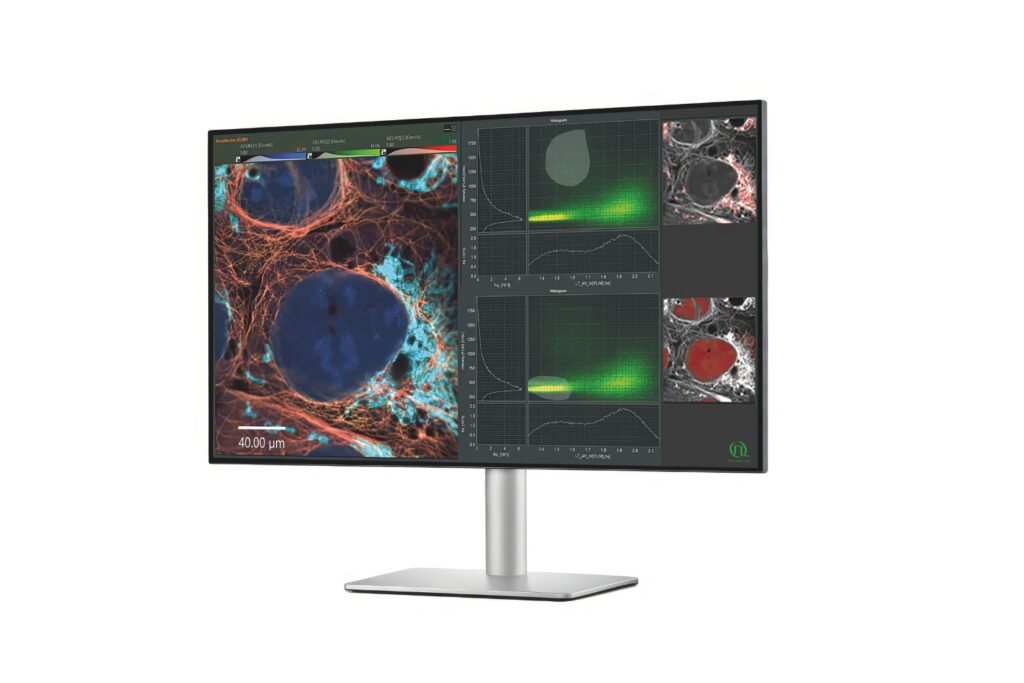
The experiments were performed using a configuration corresponding to the LSM Upgrade Kit, which adds time-resolved fluorescence capabilities to existing laser scanning microscopes. In this study, the kit was integrated into an Olympus FV3000 confocal platform and configured for quantitative FLIM-FRET measurements.
The setup included the following core components:
- A time tagging & TCSPC unit providing picosecond timing precision for fluorescence lifetime acquisition
- PMA Hybrid-40 detector enabling single-photon counting with high timing accuracy
- LDH-D-C 440 picosecond pulsed diode laser delivering synchronized excitation for decay reconstruction
- SymPhoTime 64 software for photon acquisition and multi-exponential lifetime analysis
Together, this configuration represents the detection and timing architecture required for quantitative TCSPC-based FLIM-FRET on a confocal laser scanning microscope.