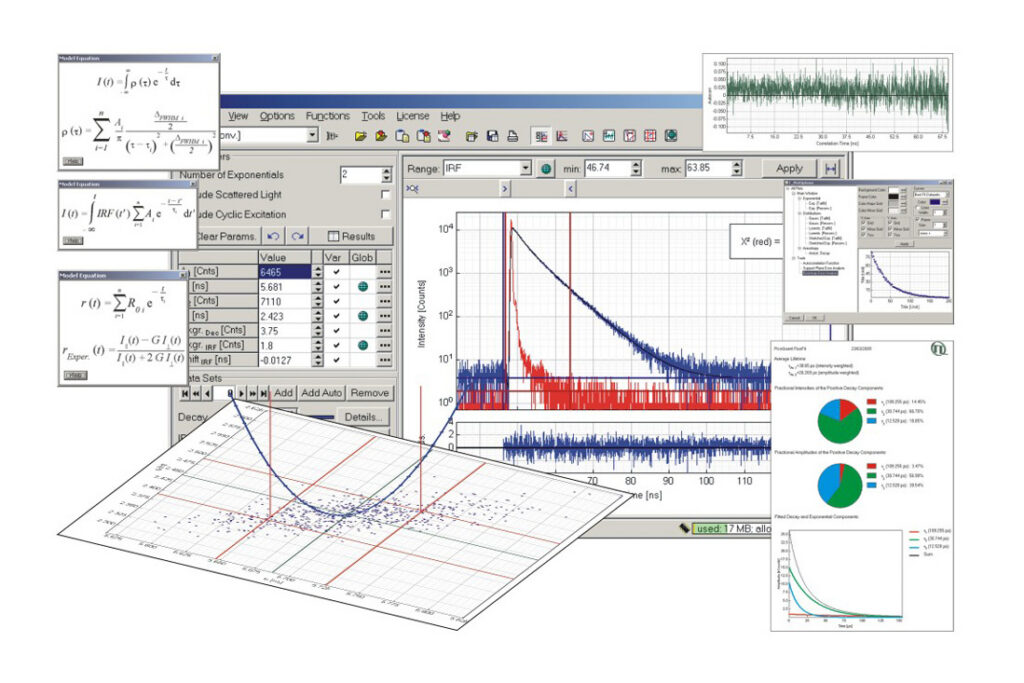
FluoFit Global Fluorescence Decay Data Analysis Software has been discontinued as of July 2018. This product is no longer available for new orders. Users looking for a comparable and updated solution are encouraged to consider its successor, the EasyTau 2 software package.
Explore the EasyTau 2 product page for detailed specifications and features.